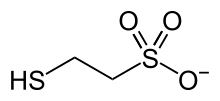
In organic chemistry, a sulfide or thioether is an organosulfur functional group with the connectivity R−S−R' as shown on right. Like many other sulfur-containing compounds, volatile sulfides have foul odors. A sulfide is similar to an ether except that it contains a sulfur atom in place of the oxygen. The grouping of oxygen and sulfur in the periodic table suggests that the chemical properties of ethers and sulfides are somewhat similar, though the extent to which this is true in practice varies depending on the application.

In organic chemistry, an epoxide is a cyclic ether, where the ether forms a three-atom ring: two atoms of carbon and one atom of oxygen. This triangular structure has substantial ring strain, making epoxides highly reactive, more so than other ethers. They are produced on a large scale for many applications. In general, low molecular weight epoxides are colourless and nonpolar, and often volatile.

A cofactor is a non-protein chemical compound or metallic ion that is required for an enzyme's role as a catalyst. Cofactors can be considered "helper molecules" that assist in biochemical transformations. The rates at which these happen are characterized in an area of study called enzyme kinetics. Cofactors typically differ from ligands in that they often derive their function by remaining bound.
Methanogens are microorganisms that produce methane as a metabolic byproduct in hypoxic conditions. They belong to the domain Archaea and are members of the phylum Euryarchaeota. Methanogens are common in wetlands, where they are responsible for marsh gas, and can occur in the digestive tracts of animals including ruminants and humans, where they are responsible for the methane content of belching and flatulence. In marine sediments, the biological production of methane, termed methanogenesis, is generally confined to where sulfates are depleted below the top layers and methanogens play an indispensable role in anaerobic wastewater treatments. Other methanogens are extremophiles, found in environments such as hot springs and submarine hydrothermal vents as well as in the "solid" rock of Earth's crust, kilometers below the surface in the deep biosphere.
Methanogenesis or biomethanation is the formation of methane coupled to energy conservation by microbes known as methanogens. Organisms capable of producing methane for energy conservation have been identified only from the domain Archaea, a group phylogenetically distinct from both eukaryotes and bacteria, although many live in close association with anaerobic bacteria. The production of methane is an important and widespread form of microbial metabolism. In anoxic environments, it is the final step in the decomposition of biomass. Methanogenesis is responsible for significant amounts of natural gas accumulations, the remainder being thermogenic.

Pterin is a heterocyclic compound composed of a pteridine ring system, with a "keto group" and an amino group on positions 4 and 2 respectively. It is structurally related to the parent bicyclic heterocycle called pteridine. Pterins, as a group, are compounds related to pterin with additional substituents. Pterin itself is of no biological significance.
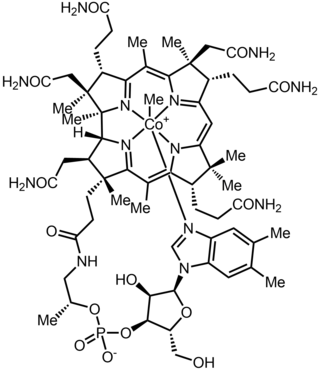
Methylcobalamin (mecobalamin, MeCbl, or MeB12) is a cobalamin, a form of vitamin B12. It differs from cyanocobalamin in that the cyano group at the cobalt is replaced with a methyl group. Methylcobalamin features an octahedral cobalt(III) centre and can be obtained as bright red crystals. From the perspective of coordination chemistry, methylcobalamin is notable as a rare example of a compound that contains metal–alkyl bonds. Nickel–methyl intermediates have been proposed for the final step of methanogenesis.
Microbial metabolism is the means by which a microbe obtains the energy and nutrients it needs to live and reproduce. Microbes use many different types of metabolic strategies and species can often be differentiated from each other based on metabolic characteristics. The specific metabolic properties of a microbe are the major factors in determining that microbe's ecological niche, and often allow for that microbe to be useful in industrial processes or responsible for biogeochemical cycles.
Tetrahydromethanopterin is a coenzyme in methanogenesis. It is the carrier of the C1 group as it is reduced to the methyl level, before transferring to the coenzyme M.
Coenzyme B is a coenzyme required for redox reactions in methanogens. The full chemical name of coenzyme B is 7-mercaptoheptanoylthreoninephosphate. The molecule contains a thiol, which is its principal site of reaction.

Methanofurans (MFRs) are a family of chemical compounds found in methanogenic archaea. These species feature a 2-aminomethylfuran linked to phenoxy group. At least three different end groups are recognized: R = tricarboxyheptanoyl (methanofuran), glutamyl-glutamyl, tricarboxy-2-hydroxyheptanoyl.

The Wood–Ljungdahl pathway is a set of biochemical reactions used by some bacteria. It is also known as the reductive acetyl-coenzyme A (Acetyl-CoA) pathway. This pathway enables these organisms to use hydrogen as an electron donor, and carbon dioxide as an electron acceptor and as a building block for biosynthesis.
In enzymology, a 2-oxopropyl-CoM reductase (carboxylating) (EC 1.8.1.5) is an enzyme that catalyzes the chemical reaction
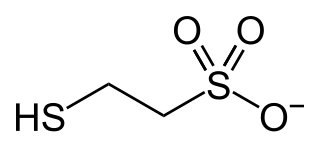
In enzymology, coenzyme-B sulfoethylthiotransferase, also known as methyl-coenzyme M reductase (MCR) or most systematically as 2-(methylthio)ethanesulfonate:N-(7-thioheptanoyl)-3-O-phosphothreonine S-(2-sulfoethyl)thiotransferase is an enzyme that catalyzes the final step in the formation of methane. It does so by combining the hydrogen donor coenzyme B and the methyl donor coenzyme M. Via this enzyme, most of the natural gas on earth was produced. Ruminants produce methane because their rumens contain methanogenic prokaryotes (Archaea) that encode and express the set of genes of this enzymatic complex.

F430 is the cofactor (sometimes called the coenzyme) of the enzyme methyl coenzyme M reductase (MCR). MCR catalyzes the reaction EC 2.8.4.1 that releases methane in the final step of methanogenesis:
Methanococcus maripaludis is a species of methanogenic archaea found in marine environments, predominantly salt marshes. M. maripaludis is a weakly motile, non-spore-forming, Gram-negative, strict anaerobic mesophile with a pleomorphic coccoid-rod shape, averaging 1.2 by 1.6 μm is size. The genome of M. maripaludis has been sequenced, and over 1,700 protein-coding genes have been identified. In ideal conditions, M. maripaludis grows quickly and can double every two hours.
The sulfate-methane transition zone (SMTZ) is a zone in oceans, lakes, and rivers typically found below the sediment surface in which sulfate and methane coexist. The formation of a SMTZ is driven by the diffusion of sulfate down the sediment column and the diffusion of methane up the sediments. At the SMTZ, their diffusion profiles meet and sulfate and methane react with one another, which allows the SMTZ to harbor a unique microbial community whose main form of metabolism is anaerobic oxidation of methane (AOM). The presence of AOM marks the transition from dissimilatory sulfate reduction to methanogenesis as the main metabolism utilized by organisms.

C1 chemistry is the chemistry of one-carbon molecules. Although many compounds and ions contain only one carbon, stable and abundant C-1 feedstocks are the focus of research. Four compounds are of major industrial importance: methane, carbon monoxide, carbon dioxide, and methanol. Technologies that interconvert these species are often used massively to match supply to demand.
In enzymology, a formylmethanofuran dehydrogenase (EC 1.2.99.5) is an enzyme that catalyzes the chemical reaction:
Ralph Stoner Wolfe was an American microbiologist, who contributed to the discovery of the single-celled archaea as the third domain of life. He was a pioneer in the biochemistry of methanogenesis.