
In genetics, complementary DNA (cDNA) is DNA synthesized from a single-stranded RNA template in a reaction catalyzed by the enzyme reverse transcriptase. cDNA is often used to express a specific protein in a cell that does not normally express that protein, or to sequence or quantify mRNA molecules using DNA based methods. cDNA that codes for a specific protein can be transferred to a recipient cell for expression, often bacterial or yeast expression systems. cDNA is also generated to analyze transcriptomic profiles in bulk tissue, single cells, or single nuclei in assays such as microarrays, qPCR, and RNA-seq.

In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part of biological inheritance. This is essential for cell division during growth and repair of damaged tissues, while it also ensures that each of the new cells receives its own copy of the DNA. The cell possesses the distinctive property of division, which makes replication of DNA essential.

The polymerase chain reaction (PCR) is a method widely used to make millions to billions of copies of a specific DNA sample rapidly, allowing scientists to amplify a very small sample of DNA sufficiently to enable detailed study. PCR was invented in 1983 by American biochemist Kary Mullis at Cetus Corporation. Mullis and biochemist Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993.

Transcription is the process of copying a segment of DNA into RNA. The segments of DNA transcribed into RNA molecules that can encode proteins produce messenger RNA (mRNA). Other segments of DNA are transcribed into RNA molecules called non-coding RNAs (ncRNAs).
A cDNA library is a combination of cloned cDNA fragments inserted into a collection of host cells, which constitute some portion of the transcriptome of the organism and are stored as a "library". cDNA is produced from fully transcribed mRNA found in the nucleus and therefore contains only the expressed genes of an organism. Similarly, tissue-specific cDNA libraries can be produced. In eukaryotic cells the mature mRNA is already spliced, hence the cDNA produced lacks introns and can be readily expressed in a bacterial cell. While information in cDNA libraries is a powerful and useful tool since gene products are easily identified, the libraries lack information about enhancers, introns, and other regulatory elements found in a genomic DNA library.
A cosmid is a type of hybrid plasmid that contains a Lambda phage cos sequence. They are often used as a cloning vector in genetic engineering. Cosmids can be used to build genomic libraries. They were first described by Collins and Hohn in 1978. Cosmids can contain 37 to 52 kb of DNA, limits based on the normal bacteriophage packaging size. They can replicate as plasmids if they have a suitable origin of replication (ori): for example SV40 ori in mammalian cells, ColE1 ori for double-stranded DNA replication, or f1 ori for single-stranded DNA replication in prokaryotes. They frequently also contain a gene for selection such as antibiotic resistance, so that the transformed cells can be identified by plating on a medium containing the antibiotic. Those cells which did not take up the cosmid would be unable to grow.

Antisense RNA (asRNA), also referred to as antisense transcript, natural antisense transcript (NAT) or antisense oligonucleotide, is a single stranded RNA that is complementary to a protein coding messenger RNA (mRNA) with which it hybridizes, and thereby blocks its translation into protein. The asRNAs have been found in both prokaryotes and eukaryotes, and can be classified into short and long non-coding RNAs (ncRNAs). The primary function of asRNA is regulating gene expression. asRNAs may also be produced synthetically and have found wide spread use as research tools for gene knockdown. They may also have therapeutic applications.
This is a list of topics in molecular biology. See also index of biochemistry articles.
A DNA construct is an artificially-designed segment of DNA borne on a vector that can be used to incorporate genetic material into a target tissue or cell. A DNA construct contains a DNA insert, called a transgene, delivered via a transformation vector which allows the insert sequence to be replicated and/or expressed in the target cell. This gene can be cloned from a naturally occurring gene, or synthetically constructed. The vector can be delivered using physical, chemical or viral methods. Typically, the vectors used in DNA constructs contain an origin of replication, a multiple cloning site, and a selectable marker. Certain vectors can carry additional regulatory elements based on the expression system involved.

A tumour inducing (Ti) plasmid is a plasmid found in pathogenic species of Agrobacterium, including A. tumefaciens, A. rhizogenes, A. rubi and A. vitis.
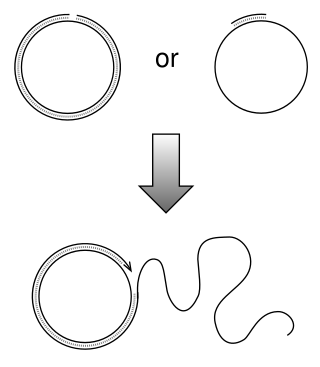

Rolling circle replication (RCR) is a process of unidirectional nucleic acid replication that can rapidly synthesize multiple copies of circular molecules of DNA or RNA, such as plasmids, the genomes of bacteriophages, and the circular RNA genome of viroids. Some eukaryotic viruses also replicate their DNA or RNA via the rolling circle mechanism.
In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complement in specifying a sequence of amino acids. Depending on the context, sense may have slightly different meanings. For example, the negative-sense strand of DNA is equivalent to the template strand, whereas the positive-sense strand is the non-template strand whose nucleotide sequence is equivalent to the sequence of the mRNA transcript.
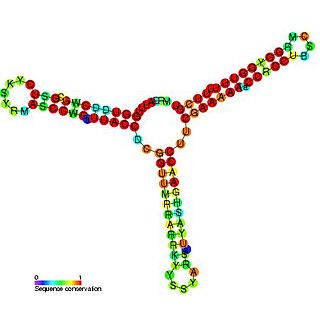
RNAI is a non-coding RNA that is an antisense repressor of the replication of some E. coli plasmids, including ColE1. Plasmid replication is usually initiated by RNAII, which acts as a primer by binding to its template DNA. The complementary RNAI binds RNAII prohibiting it from its initiation role. The rate of degradation of RNAI is therefore a major factor in the control of plasmid replication. This rate of degradation is aided by the pcnB gene product, which polyadenylates the 3' end of RNAI targeting it for degradation by PNPase.
In molecular cloning, a vector is any particle used as a vehicle to artificially carry a foreign nucleic sequence – usually DNA – into another cell, where it can be replicated and/or expressed. A vector containing foreign DNA is termed recombinant DNA. The four major types of vectors are plasmids, viral vectors, cosmids, and artificial chromosomes. Of these, the most commonly used vectors are plasmids. Common to all engineered vectors are an origin of replication, a multicloning site, and a selectable marker.
The versatility of polymerase chain reaction (PCR) has led to modifications of the basic protocol being used in a large number of variant techniques designed for various purposes. This article summarizes many of the most common variations currently or formerly used in molecular biology laboratories; familiarity with the fundamental premise by which PCR works and corresponding terms and concepts is necessary for understanding these variant techniques.
Φ29 DNA polymerase is an enzyme from the bacteriophage Φ29. It is being increasingly used in molecular biology for multiple displacement DNA amplification procedures, and has a number of features that make it particularly suitable for this application. It was discovered and characterized by Spanish scientists Luis Blanco and Margarita Salas.
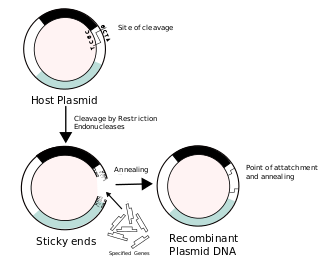
Recombinant DNA (rDNA), or molecular cloning, is the process by which a single gene, or segment of DNA, is isolated and amplified. Recombinant DNA is also known as in vitro recombination. A cloning vector is a DNA molecule that carries foreign DNA into a host cell, where it replicates, producing many copies of itself along with the foreign DNA. There are many types of cloning vectors such as plasmids and phages. In order to carry out recombination between vector and the foreign DNA, it is necessary the vector and DNA to be cloned by digestion, ligase the foreign DNA into the vector with the enzyme DNA ligase. And DNA is inserted by introducing the DNA into bacteria cells by transformation.

An R-loop is a three-stranded nucleic acid structure, composed of a DNA:RNA hybrid and the associated non-template single-stranded DNA. R-loops may be formed in a variety of circumstances and may be tolerated or cleared by cellular components. The term "R-loop" was given to reflect the similarity of these structures to D-loops; the "R" in this case represents the involvement of an RNA moiety.
In cellular biology, the plasmid copy number is the number of copies of a given plasmid in a cell. To ensure survival and thus the continued propagation of the plasmid, they must regulate their copy number. If a plasmid has too high of a copy number, they may excessively burden their host by occupying too much cellular machinery and using too much energy. On the other hand, too low of a copy number may result in the plasmid not being present in all of their host's progeny. Plasmids may be either low, medium or high copy number plasmids; the regulation mechanisms between low and medium copy number plasmids are different. Low copy plasmids require either a partitioning system or a toxin-antitoxin pair such as CcdA/CcdB to ensure that each daughter receives the plasmid. For example, the F plasmid, which is the origin of BACs is a single copy plasmid with a partitioning system encoded in an operon right next to the plasmid origin. The partitioning system interacts with the septation apparatus to ensure that each daughter receives a copy of the plasmid. Many biotechnology applications utilize mutated plasmids that replicate to high copy number. For example, pBR322 is a medium copy number plasmid from which several high copy number cloning vectors have been derived by mutagenesis, such as the well known pUC series. This delivers the convenience of high plasmid DNA yields but the additional burden of the high copy number restricts the plasmid size. Larger high copy plasmids (>30kb) are disfavoured and also prone to size reduction through deletional mutagenesis.
This glossary of genetics is a list of definitions of terms and concepts commonly used in the study of genetics and related disciplines in biology, including molecular biology, cell biology, and evolutionary biology. It is intended as introductory material for novices; for more specific and technical detail, see the article corresponding to each term. For related terms, see Glossary of evolutionary biology.










