Entamoeba is a genus of Amoebozoa found as internal parasites or commensals of animals. In 1875, Fedor Lösch described the first proven case of amoebic dysentery in St. Petersburg, Russia. He referred to the amoeba he observed microscopically as Amoeba coli; however, it is not clear whether he was using this as a descriptive term or intended it as a formal taxonomic name. The genus Entamoeba was defined by Casagrandi and Barbagallo for the species Entamoeba coli, which is known to be a commensal organism. Lösch's organism was renamed Entamoeba histolytica by Fritz Schaudinn in 1903; he later died, in 1906, from a self-inflicted infection when studying this amoeba. For a time during the first half of the 20th century the entire genus Entamoeba was transferred to Endamoeba, a genus of amoebas infecting invertebrates about which little is known. This move was reversed by the International Commission on Zoological Nomenclature in the late 1950s, and Entamoeba has stayed 'stable' ever since.

Meiosis is a special type of cell division of germ cells and apicomplexans in sexually-reproducing organisms that produces the gametes, such as sperm or egg cells. It involves two rounds of division that ultimately result in four cells with only one copy of each chromosome (haploid). Additionally, prior to the division, genetic material from the paternal and maternal copies of each chromosome is crossed over, creating new combinations of code on each chromosome. Later on, during fertilisation, the haploid cells produced by meiosis from a male and a female will fuse to create a cell with two copies of each chromosome again, the zygote.

Chromosomal crossover, or crossing over, is the exchange of genetic material during sexual reproduction between two homologous chromosomes' non-sister chromatids that results in recombinant chromosomes. It is one of the final phases of genetic recombination, which occurs in the pachytene stage of prophase I of meiosis during a process called synapsis. Synapsis begins before the synaptonemal complex develops and is not completed until near the end of prophase I. Crossover usually occurs when matching regions on matching chromosomes break and then reconnect to the other chromosome.

Genetic recombination is the exchange of genetic material between different organisms which leads to production of offspring with combinations of traits that differ from those found in either parent. In eukaryotes, genetic recombination during meiosis can lead to a novel set of genetic information that can be further passed on from parents to offspring. Most recombination occurs naturally and can be classified into two types: (1) interchromosomal recombination, occurring through independent assortment of alleles whose loci are on different but homologous chromosomes ; & (2) intrachromosomal recombination, occurring through crossing over.

A couple of homologous chromosomes, or homologs, are a set of one maternal and one paternal chromosome that pair up with each other inside a cell during fertilization. Homologs have the same genes in the same loci, where they provide points along each chromosome that enable a pair of chromosomes to align correctly with each other before separating during meiosis. This is the basis for Mendelian inheritance, which characterizes inheritance patterns of genetic material from an organism to its offspring parent developmental cell at the given time and area.
RecQ helicase is a family of helicase enzymes initially found in Escherichia coli that has been shown to be important in genome maintenance. They function through catalyzing the reaction ATP + H2O → ADP + P and thus driving the unwinding of paired DNA and translocating in the 3' to 5' direction. These enzymes can also drive the reaction NTP + H2O → NDP + P to drive the unwinding of either DNA or RNA.

RecA is a 38 kilodalton protein essential for the repair and maintenance of DNA. A RecA structural and functional homolog has been found in every species in which one has been seriously sought and serves as an archetype for this class of homologous DNA repair proteins. The homologous protein is called RAD51 in eukaryotes and RadA in archaea.
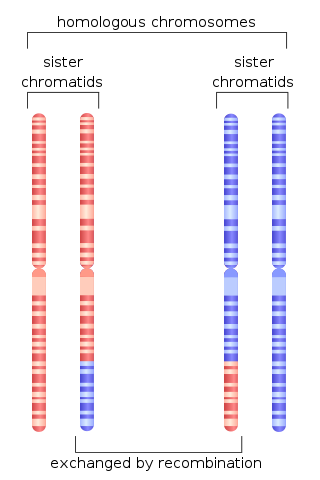
Homologous recombination is a type of genetic recombination in which genetic information is exchanged between two similar or identical molecules of double-stranded or single-stranded nucleic acids.
Recombinases are genetic recombination enzymes.
DNA repair protein RAD51 homolog 1 is a protein encoded by the gene RAD51. The enzyme encoded by this gene is a member of the RAD51 protein family which assists in repair of DNA double strand breaks. RAD51 family members are homologous to the bacterial RecA, Archaeal RadA and yeast Rad51. The protein is highly conserved in most eukaryotes, from yeast to humans.

Bloom syndrome protein is a protein that in humans is encoded by the BLM gene and is not expressed in Bloom syndrome.

RAD52 homolog , also known as RAD52, is a protein which in humans is encoded by the RAD52 gene.

DNA repair and recombination protein RAD54-like is a protein that in humans is encoded by the RAD54L gene.
In molecular biology, a displacement loop or D-loop is a DNA structure where the two strands of a double-stranded DNA molecule are separated for a stretch and held apart by a third strand of DNA. An R-loop is similar to a D-loop, but in this case the third strand is RNA rather than DNA. The third strand has a base sequence which is complementary to one of the main strands and pairs with it, thus displacing the other complementary main strand in the region. Within that region the structure is thus a form of triple-stranded DNA. A diagram in the paper introducing the term illustrated the D-loop with a shape resembling a capital "D", where the displaced strand formed the loop of the "D".

MutS protein homolog 4 is a protein that in humans is encoded by the MSH4 gene.

DNA repair and recombination protein RAD54B is a protein that in humans is encoded by the RAD54B gene.

DNA replication licensing factor MCM8 is a protein that in humans is encoded by the MCM8 gene.

The meiotic recombination checkpoint monitors meiotic recombination during meiosis, and blocks the entry into metaphase I if recombination is not efficiently processed.

Synthesis-dependent strand annealing (SDSA) is a major mechanism of homology-directed repair of DNA double-strand breaks (DSBs). Although many of the features of SDSA were first suggested in 1976, the double-Holliday junction model proposed in 1983 was favored by many researchers. In 1994, studies of double-strand gap repair in Drosophila were found to be incompatible with the double-Holliday junction model, leading researchers to propose a model they called synthesis-dependent strand annealing. Subsequent studies of meiotic recombination in S. cerevisiae found that non-crossover products appear earlier than double-Holliday junctions or crossover products, challenging the previous notion that both crossover and non-crossover products are produced by double-Holliday junctions and leading the authors to propose that non-crossover products are generated through SDSA.
GT198 is a human oncogene located within the BRCA1 locus at chromosome 17q21. It encodes protein product named GT198, Hop2 or TBPIP. The GT198 gene is found to be mutated with its protein overexpressed in human cancers including breast and ovarian cancers.