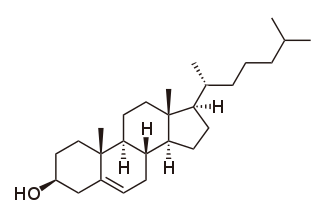
Cholesterol is the principal sterol of all higher animals, distributed in body tissues, especially the brain and spinal cord, and in animal fats and oils.

Lipid-anchored proteins are proteins located on the surface of the cell membrane that are covalently attached to lipids embedded within the cell membrane. These proteins insert and assume a place in the bilayer structure of the membrane alongside the similar fatty acid tails. The lipid-anchored protein can be located on either side of the cell membrane. Thus, the lipid serves to anchor the protein to the cell membrane. They are a type of proteolipids.

A general anaesthetic is a drug that brings about a reversible loss of consciousness. These drugs are generally administered by an anaesthetist/anesthesiologist to induce or maintain general anaesthesia to facilitate surgery.

The plasma membranes of cells contain combinations of glycosphingolipids, cholesterol and protein receptors organised in glycolipoprotein lipid microdomains termed lipid rafts. Their existence in cellular membranes remains controversial. Indeed, Kervin and Overduin imply that lipid rafts are misconstrued protein islands, which they propose form through a proteolipid code. Nonetheless, it has been proposed that they are specialized membrane microdomains which compartmentalize cellular processes by serving as organising centers for the assembly of signaling molecules, allowing a closer interaction of protein receptors and their effectors to promote kinetically favorable interactions necessary for the signal transduction. Lipid rafts influence membrane fluidity and membrane protein trafficking, thereby regulating neurotransmission and receptor trafficking. Lipid rafts are more ordered and tightly packed than the surrounding bilayer, but float freely within the membrane bilayer. Although more common in the cell membrane, lipid rafts have also been reported in other parts of the cell, such as the Golgi apparatus and lysosomes.
Phosphatidic acids are anionic phospholipids important to cell signaling and direct activation of lipid-gated ion channels. Hydrolysis of phosphatidic acid gives rise to one molecule each of glycerol and phosphoric acid and two molecules of fatty acids. They constitute about 0.25% of phospholipids in the bilayer.

Myristoylation is a lipidation modification where a myristoyl group, derived from myristic acid, is covalently attached by an amide bond to the alpha-amino group of an N-terminal glycine residue. Myristic acid is a 14-carbon saturated fatty acid (14:0) with the systematic name of n-tetradecanoic acid. This modification can be added either co-translationally or post-translationally. N-myristoyltransferase (NMT) catalyzes the myristic acid addition reaction in the cytoplasm of cells. This lipidation event is the most common type of fatty acylation and is present in many organisms, including animals, plants, fungi, protozoans and viruses. Myristoylation allows for weak protein–protein and protein–lipid interactions and plays an essential role in membrane targeting, protein–protein interactions and functions widely in a variety of signal transduction pathways.

Phosphoinositide phospholipase C is a family of eukaryotic intracellular enzymes that play an important role in signal transduction processes. These enzymes belong to a larger superfamily of Phospholipase C. Other families of phospholipase C enzymes have been identified in bacteria and trypanosomes. Phospholipases C are phosphodiesterases.
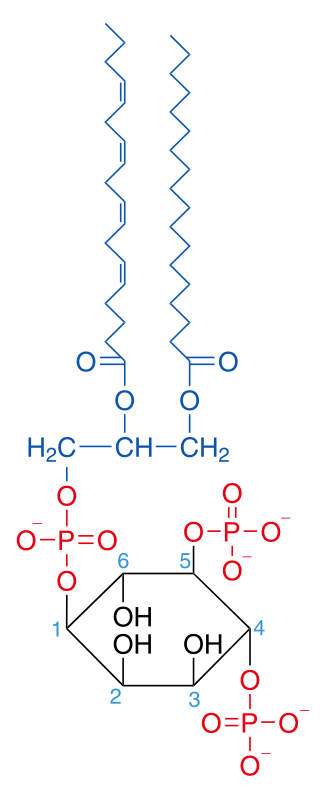
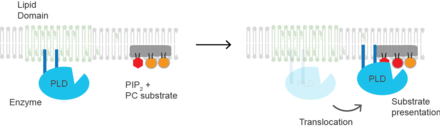
Phosphatidylinositol 4,5-bisphosphate or PtdIns(4,5)P2, also known simply as PIP2 or PI(4,5)P2, is a minor phospholipid component of cell membranes. PtdIns(4,5)P2 is enriched at the plasma membrane where it is a substrate for a number of important signaling proteins. PIP2 also forms lipid clusters that sort proteins.

Lipid signaling, broadly defined, refers to any biological cell signaling event involving a lipid messenger that binds a protein target, such as a receptor, kinase or phosphatase, which in turn mediate the effects of these lipids on specific cellular responses. Lipid signaling is thought to be qualitatively different from other classical signaling paradigms because lipids can freely diffuse through membranes. One consequence of this is that lipid messengers cannot be stored in vesicles prior to release and so are often biosynthesized "on demand" at their intended site of action. As such, many lipid signaling molecules cannot circulate freely in solution but, rather, exist bound to special carrier proteins in serum.
Phospholipase D (EC 3.1.4.4, lipophosphodiesterase II, lecithinase D, choline phosphatase, PLD; systematic name phosphatidylcholine phosphatidohydrolase) is an enzyme of the phospholipase superfamily that catalyses the following reaction
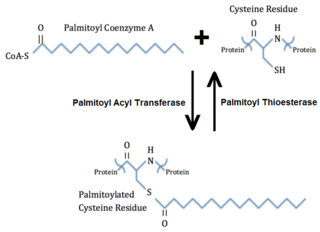
Palmitoylation is the covalent attachment of fatty acids, such as palmitic acid, to cysteine (S-palmitoylation) and less frequently to serine and threonine (O-palmitoylation) residues of proteins, which are typically membrane proteins. The precise function of palmitoylation depends on the particular protein being considered. Palmitoylation enhances the hydrophobicity of proteins and contributes to their membrane association. Palmitoylation also appears to play a significant role in subcellular trafficking of proteins between membrane compartments, as well as in modulating protein–protein interactions. In contrast to prenylation and myristoylation, palmitoylation is usually reversible (because the bond between palmitic acid and protein is often a thioester bond). The reverse reaction in mammalian cells is catalyzed by acyl-protein thioesterases (APTs) in the cytosol and palmitoyl protein thioesterases in lysosomes. Because palmitoylation is a dynamic, post-translational process, it is believed to be employed by the cell to alter the subcellular localization, protein–protein interactions, or binding capacities of a protein.

Phospholipase D2 is an enzyme that in humans is encoded by the PLD2 gene.

Phospholipase C (PLC) is a class of membrane-associated enzymes that cleave phospholipids just before the phosphate group (see figure). It is most commonly taken to be synonymous with the human forms of this enzyme, which play an important role in eukaryotic cell physiology, in particular signal transduction pathways. Phospholipase C's role in signal transduction is its cleavage of phosphatidylinositol 4,5-bisphosphate (PIP2) into diacyl glycerol (DAG) and inositol 1,4,5-trisphosphate (IP3), which serve as second messengers. Activators of each PLC vary, but typically include heterotrimeric G protein subunits, protein tyrosine kinases, small G proteins, Ca2+, and phospholipids.

Potassium channel subfamily K member 2, also known as TREK-1, is a protein that in humans is encoded by the KCNK2 gene.
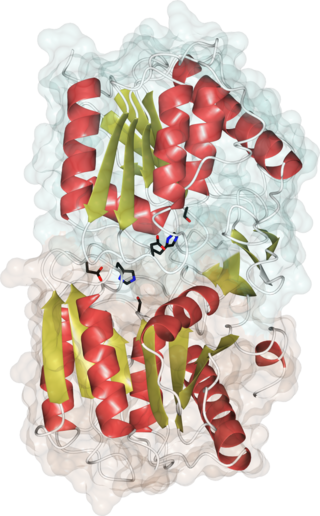
Acyl-protein thioesterases are enzymes that cleave off lipid modifications on proteins, located on the sulfur atom of cysteine residues linked via a thioester bond. Acyl-protein thioesterases are part of the α/β hydrolase superfamily of proteins and have a conserved catalytic triad. For that reason, acyl-protein thioesterases are also able to hydrolyze oxygen-linked ester bonds.
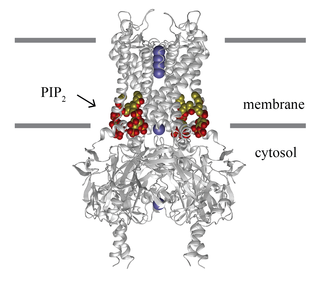
Lipid-gated ion channels are a class of ion channels whose conductance of ions through the membrane depends directly on lipids. Classically the lipids are membrane resident anionic signaling lipids that bind to the transmembrane domain on the inner leaflet of the plasma membrane with properties of a classic ligand. Other classes of lipid-gated channels include the mechanosensitive ion channels that respond to lipid tension, thickness, and hydrophobic mismatch. A lipid ligand differs from a lipid cofactor in that a ligand derives its function by dissociating from the channel while a cofactor typically derives its function by remaining bound.
Palmitate mediated localization is a biological process that trafficks a palmitoylated protein to ordered lipid domains.

Cholesterol is a cell signaling molecule that is highly regulated in eukaryotic cell membranes. In human health, its effects are most notable in inflammation, metabolic syndrome, and neurodegeneration. At the molecular level, cholesterol primarily signals by regulating clustering of saturated lipids and proteins that depend on clustering for their regulation.
PIP2 domains are a type of cholesterol-independent lipid domain formed from phosphatidylinositol and positively charged proteins in the plasma membrane. They tend to inhibit GM1 lipid raft function.
Membrane-mediated anesthesia or anaesthesia (UK) is a mechanism of action that involves an anesthetic exerting its effects through the lipid membrane. Established mechanism exists for both general and local anesthetics. The anesthetic binding site is within ordered lipids and binding disrupts the function of the ordered lipid. See Theories of general anaesthetic action for a broader discussion of purely theoretical mechanisms.