Related Research Articles

The human genome is a complete set of nucleic acid sequences for humans, encoded as DNA within the 23 chromosome pairs in cell nuclei and in a small DNA molecule found within individual mitochondria. These are usually treated separately as the nuclear genome and the mitochondrial genome. Human genomes include both protein-coding DNA sequences and various types of DNA that does not encode proteins. The latter is a diverse category that includes DNA coding for non-translated RNA, such as that for ribosomal RNA, transfer RNA, ribozymes, small nuclear RNAs, and several types of regulatory RNAs. It also includes promoters and their associated gene-regulatory elements, DNA playing structural and replicatory roles, such as scaffolding regions, telomeres, centromeres, and origins of replication, plus large numbers of transposable elements, inserted viral DNA, non-functional pseudogenes and simple, highly repetitive sequences. Introns make up a large percentage of non-coding DNA. Some of this non-coding DNA is non-functional junk DNA, such as pseudogenes, but there is no firm consensus on the total amount of junk DNA.
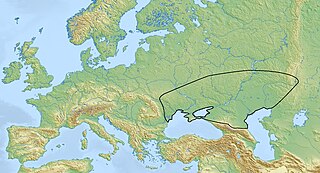
The Yamnaya culture or the Yamna culture, also known as the Pit Grave culture or Ochre Grave culture, was a late Copper Age to early Bronze Age archaeological culture of the region between the Southern Bug, Dniester, and Ural rivers, dating to 3300–2600 BCE. It was discovered by Vasily Gorodtsov following his archaeological excavations near the Donets River in 1901–1903. Its name derives from its characteristic burial tradition: Я́мная is a Russian adjective that means 'related to pits ', as these people used to bury their dead in tumuli (kurgans) containing simple pit chambers.
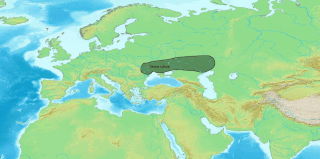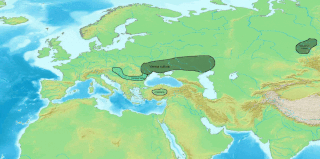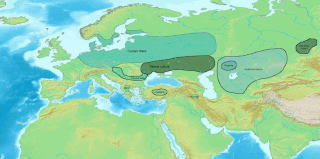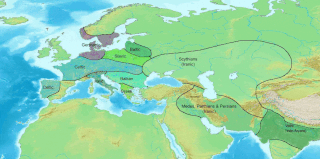
The Indo-Aryan migrations were the migrations into the Indian subcontinent of Indo-Aryan peoples, an ethnolinguistic group that spoke Indo-Aryan languages, the predominant languages of today's North India, Pakistan, Nepal, Bangladesh, Sri Lanka and the Maldives. Indo-Aryan population movements into the region from Central Asia are considered to have started after 2000 BCE, as a slow diffusion during the Late Harappan period, which led to a language shift in the northern Indian subcontinent. Several hundred years later, the Iranian languages were brought into the Iranian plateau by the Iranians, who were closely related to the Indo-Aryans.

The Chimpanzee Genome Project was an effort to determine the DNA sequence of the chimpanzee genome. Sequencing began in 2005 and by 2013 twenty-four individual chimpanzees had been sequenced. This project was folded into the Great Ape Genome Project.
Genetics and archaeogenetics of South Asia is the study of the genetics and archaeogenetics of the ethnic groups of South Asia. It aims at uncovering these groups' genetic histories. The geographic position of the Indian subcontinent makes its biodiversity important for the study of the early dispersal of anatomically modern humans across Asia.

Human genetic variation is the genetic differences in and among populations. There may be multiple variants of any given gene in the human population (alleles), a situation called polymorphism.
Human evolutionary genetics studies how one human genome differs from another human genome, the evolutionary past that gave rise to the human genome, and its current effects. Differences between genomes have anthropological, medical, historical and forensic implications and applications. Genetic data can provide important insights into human evolution.
The Karitiana or Caritiana are an indigenous people of Brazil, whose reservation is located in the western Amazon. They count 320 members, and the leader of their tribal association is Renato Caritiana. They subsist by farming, fishing and hunting, and have almost no contact with the outside world. Their tongue, the Karitiâna language, is an Arikém language of Brazil.
The chimpanzee–human last common ancestor (CHLCA) is the last common ancestor shared by the extant Homo (human) and Pan genera of Hominini. Estimates of the divergence date vary widely from thirteen to five million years ago.

In paleoanthropology, the recent African origin of modern humans or the "Out of Africa" theory (OOA) is the most widely accepted model of the geographic origin and early migration of anatomically modern humans. It follows the early expansions of hominins out of Africa, accomplished by Homo erectus and then Homo neanderthalensis.

The peopling of India refers to the migration of Homo sapiens into the Indian subcontinent. Anatomically modern humans settled India in multiple waves of early migrations, over tens of millennia. The first migrants came with the Coastal Migration/Southern Dispersal 65,000 years ago, whereafter complex migrations within south and southeast Asia took place. West-Asian (Iranian) hunter-gatherers migrated to South Asia after the Last Glacial Period but before the onset of farming. Together with ancient South Asian hunter-gatherers they formed the population of the Indus Valley civilisation (IVC).
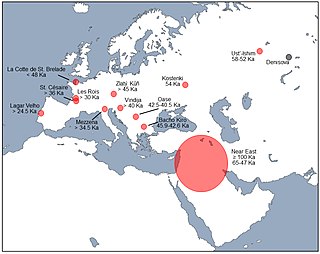
Interbreeding between archaic and modern humans occurred during the Middle Paleolithic and early Upper Paleolithic. The interbreeding happened in several independent events that included Neanderthals and Denisovans, as well as several unidentified hominins.
Incomplete lineage sorting, also termed hemiplasy, deep coalescence, retention of ancestral polymorphism, or trans-species polymorphism, describes a phenomenon in population genetics when ancestral gene copies fail to coalesce into a common ancestral copy until deeper than previous speciation events. It is caused by lineage sorting of genetic polymorphisms that were retained across successive nodes in the species tree. In other words, the tree produced by a single gene differs from the population or species level tree, producing a discordant tree. Whatever the mechanism, the result is that a generated species level tree may differ depending on the selected genes used for assessment. This is in contrast to complete lineage sorting, where the tree produced by the gene is the same as the population or species level tree. Both are common results in phylogenetic analysis, although it depends on the gene, organism, and sampling technique.

The study of the genetics and archaeogenetics of the Gujarati people of India aims at uncovering these people's genetic history. According to the 1000 Genomes Project, "Gujarati" is a general term used to describe people who trace their ancestry to the region of Gujarat, located in the northwestern part of the Indian subcontinent, and who speak the Gujarati language, an Indo-European language. They have some genetic commonalities as well as differences with other ethnic groups of India.

In archaeogenetics, the term Ancient North Eurasian (ANE) is the name given to an ancestral component that represents the lineage of the people of the Mal'ta–Buret' culture (c. 24,000 BP) and populations closely related to them, such as the Upper Paleolithic individuals from Afontova Gora in Siberia. Genetic studies indicate that the ANE are closely related to the Ancient North Siberians (ANS) represented by two ancient specimens from the preceding Yana Culture (c. 32,000 BP). The ANE can either be considered to descend from the earlier ANS population, or that both ANE and ANS are closely related, albeit differentiated, sister lineages, with both having originated from an 'Early West Eurasian' hunter-gatherer lineage (represented by Kostenki-14, c. 40,000 BP), which absorbed an 'Early East Eurasian' population (represented by the Tianyuan man, c. 40,000 BP). The ANS and ANE each derive between 18% to one third of their ancestry from an Early East Eurasian lineage and between two thirds to 82% from an Early West Eurasian lineage.
Genetic studies on Neanderthal ancient DNA became possible in the late 1990s. The Neanderthal genome project, established in 2006, presented the first fully sequenced Neanderthal genome in 2013.

Who We Are and How We Got Here is a 2018 book on the contribution of genome-wide ancient DNA research to human population genetics by the geneticist David Reich. He describes discoveries made by his group and others, based on analysis and comparison of ancient and modern DNA from human populations around the world. Central to these is the finding that almost all human populations are mixtures resulting from multiple population migrations and gene flow.

In archaeogenetics, the term Western Hunter-Gatherer (WHG), West European Hunter-Gatherer, Western European Hunter-Gatherer, Villabruna cluster, or Oberkassel cluster names a distinct ancestral component of modern Europeans, representing descent from a population of Mesolithic hunter-gatherers who scattered over Western, Southern and Central Europe, from the British Isles in the west to the Carpathians in the east, following the retreat of the ice sheet of the Last Glacial Maximum.
Caucasus hunter-gatherer (CHG), also called Satsurblia cluster, is an anatomically modern human genetic lineage, first identified in a 2015 study, based on the population genetics of several modern Western Eurasian populations.
References
- ↑ "365 days: Nature's 10". Nature. 528 (7583): 459–467. 2015. Bibcode:2015Natur.528..459.. doi: 10.1038/528459a . ISSN 0028-0836. PMID 26701036.
- ↑ https://www.york.ac.uk/archaeology/news-and-events/news/external/news2022/antiquity-prize-win/
- 1 2 Reich, David Emile (1999). Genetic analysis of human evolutionary history with implications for gene mapping. ox.ac.uk (DPhil thesis). University of Oxford. OCLC 863264589. EThOS uk.bl.ethos.580823.

- ↑ "David Reich | Genetics". genetics.hms.harvard.edu. Retrieved 2018-01-08.
- ↑ "Nature's 10". Nature. 528 (7583): 459–467. December 2015. Bibcode:2015Natur.528..459.. doi: 10.1038/528459a . PMID 26701036. S2CID 4450003.
- ↑ Massry Prize 2021
- 1 2 3 Zimmer, Carl (2018-03-20). "David Reich Unearths Human History Etched in Bone". The New York Times. Retrieved 2018-03-20.
- ↑ Rincon, Paul (11 April 2018). "How ancient DNA is transforming our view of the past". BBC. Retrieved 11 April 2018.
- ↑ Emile., Reich, David (1999). Genetic analysis of human evolutionary history with implications for gene mapping (Thesis). University of Oxford.
{{cite thesis}}: CS1 maint: multiple names: authors list (link) - ↑ ScienceNews.org – 'Hybrid-Driven Evolution: Genomes show complexity of human-chimp split: Not only did the evolutionary parting of human from chimpanzee ancestors occur more recently than had been indicated by previous data, but it also played out over an extended period during which forerunners of people and chimps interbred', Bruce Bower, Science News (May 20, 2006)
- ↑ Patterson, N.; Richter, D. J.; Gnerre, S.; Lander, E. S.; Reich, D. (2006). "Genetic evidence for complex speciation of humans and chimpanzees". Nature. 441 (7097): 1103–1108. Bibcode:2006Natur.441.1103P. doi:10.1038/nature04789. PMID 16710306. S2CID 2325560.
- ↑ Reich 2009.
- ↑ Chakravarti, Aravinda (24 September 2009). "Tracing India's invisible lthreads" (PDF). Nature (News & Views). Archived from the original (PDF) on 3 September 2015. Retrieved 16 March 2016.
- ↑ Dolgin, Elie (September 23, 2009). "Indian ancestry revealed". Nature. doi:10.1038/news.2009.935 – via www.nature.com.
- ↑ Srinath Perur, The origins of Indians. What our genes are telling us., Fountain Ink
- ↑ Metspalu et al. 2011.
- ↑ David Cameron (July 20, 2011). "Detail distinguishes map of African-American genomics". Harvard Gazette. Retrieved July 22, 2011.
- ↑ Reich, D.; Green, R.E.; Kircher, M.; Krause, J.; Patterson, N.; Durand, E.Y.; et al. (2010). "Genetic history of an archaic hominin group from Denisova Cave in Siberia". Nature. 468 (7327): 1053–1060. Bibcode:2010Natur.468.1053R. doi:10.1038/nature09710. PMC 4306417 . PMID 21179161.Reich, D.; Patterson, N.; Kircher, M.; Delfin, F.; Nandineni, M.R.; Pugach, I.; et al. (2011). "Denisova Admixture and the First Modern Human Dispersals into Southeast Asia and Oceania". The American Journal of Human Genetics. 89 (4): 516–528. doi:10.1016/j.ajhg.2011.09.005. PMC 3188841 . PMID 21944045.Sankararaman, S.; Patterson, N.; Li, H.; Pääbo, S.; Reich, D; Akey, J.M. (2012). "The Date of Interbreeding between Neandertals and Modern Humans". PLOS Genetics. 8 (10): e1002947. arXiv: 1208.2238 . doi: 10.1371/journal.pgen.1002947 . PMC 3464203 . PMID 23055938. Carl Zimmer, "Interbreeding with Neanderthals", Discover , March 2013, pp. 38–44.
- ↑ https://reich.hms.harvard.edu/sites/reich.hms.harvard.edu/files/inline-files/2007_NG_Haiman_colorectal_and_prostate.pdf [ bare URL PDF ]
- 1 2 Who We Are and How We Got Here: Ancient DNA and the New Science of the Human Past. By David Reich. New York: Pantheon, 2018.
- ↑ "Inferring demographic history from genetic data". uqrmaie1.github.io. Retrieved 2022-05-18.