Related Research Articles
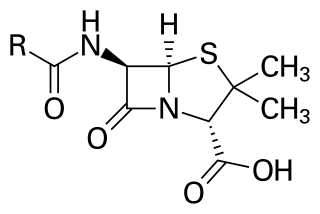
Penicillins are a group of β-lactam antibiotics originally obtained from Penicillium moulds, principally P. chrysogenum and P. rubens. Most penicillins in clinical use are synthesised by P. chrysogenum using deep tank fermentation and then purified. A number of natural penicillins have been discovered, but only two purified compounds are in clinical use: penicillin G and penicillin V. Penicillins were among the first medications to be effective against many bacterial infections caused by staphylococci and streptococci. They are still widely used today for different bacterial infections, though many types of bacteria have developed resistance following extensive use.
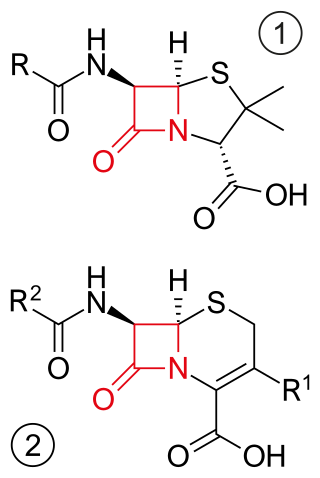
β-lactam antibiotics are antibiotics that contain a beta-lactam ring in their chemical structure. This includes penicillin derivatives (penams), cephalosporins and cephamycins (cephems), monobactams, carbapenems and carbacephems. Most β-lactam antibiotics work by inhibiting cell wall biosynthesis in the bacterial organism and are the most widely used group of antibiotics. Until 2003, when measured by sales, more than half of all commercially available antibiotics in use were β-lactam compounds. The first β-lactam antibiotic discovered, penicillin, was isolated from a strain of Penicillium rubens.

Staphylococcus aureus is a Gram-positive spherically shaped bacterium, a member of the Bacillota, and is a usual member of the microbiota of the body, frequently found in the upper respiratory tract and on the skin. It is often positive for catalase and nitrate reduction and is a facultative anaerobe that can grow without the need for oxygen. Although S. aureus usually acts as a commensal of the human microbiota, it can also become an opportunistic pathogen, being a common cause of skin infections including abscesses, respiratory infections such as sinusitis, and food poisoning. Pathogenic strains often promote infections by producing virulence factors such as potent protein toxins, and the expression of a cell-surface protein that binds and inactivates antibodies. S. aureus is one of the leading pathogens for deaths associated with antimicrobial resistance and the emergence of antibiotic-resistant strains, such as methicillin-resistant S. aureus (MRSA), is a worldwide problem in clinical medicine. Despite much research and development, no vaccine for S. aureus has been approved.

Methicillin-resistant Staphylococcus aureus (MRSA) is a group of gram-positive bacteria that are genetically distinct from other strains of Staphylococcus aureus. MRSA is responsible for several difficult-to-treat infections in humans. It caused more than 100,000 deaths worldwide attributable to antimicrobial resistance in 2019.

Methicillin (USAN), also known as meticillin (INN), is a narrow-spectrum β-lactam antibiotic of the penicillin class.

The cephalosporins are a class of β-lactam antibiotics originally derived from the fungus Acremonium, which was previously known as Cephalosporium.

Vancomycin-resistant Staphylococcus aureus (VRSA) are strains of Staphylococcus aureus that have acquired resistance to the glycopeptide antibiotic vancomycin. Bacteria can acquire resistant genes either by random mutation or through the transfer of DNA from one bacterium to another. Resistance genes interfere with the normal antibiotic function and allow a bacteria to grow in the presence of the antibiotic. Resistance in VRSA is conferred by the plasmid-mediated vanA gene and operon. Although VRSA infections are uncommon, VRSA is often resistant to other types of antibiotics and a potential threat to public health because treatment options are limited. VRSA is resistant to many of the standard drugs used to treat S. aureus infections. Furthermore, resistance can be transferred from one bacterium to another.

Staphylococcus haemolyticus is a member of the coagulase-negative staphylococci (CoNS). It is part of the skin flora of humans, and its largest populations are usually found at the axillae, perineum, and inguinal areas. S. haemolyticus also colonizes primates and domestic animals. It is a well-known opportunistic pathogen, and is the second-most frequently isolated CoNS. Infections can be localized or systemic, and are often associated with the insertion of medical devices. The highly antibiotic-resistant phenotype and ability to form biofilms make S. haemolyticus a difficult pathogen to treat. Its most closely related species is Staphylococcus borealis.

Dicloxacillin is a narrow-spectrum β-lactam antibiotic of the penicillin class. It is used to treat infections caused by susceptible (non-resistant) Gram-positive bacteria. It is active against beta-lactamase-producing organisms such as Staphylococcus aureus, which would otherwise be resistant to most penicillins. Dicloxacillin is available under a variety of trade names including Diclocil (BMS).
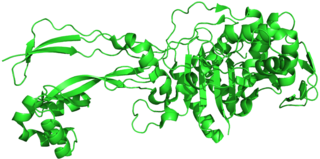
Penicillin-binding proteins (PBPs) are a group of proteins that are characterized by their affinity for and binding of penicillin. They are a normal constituent of many bacteria; the name just reflects the way by which the protein was discovered. All β-lactam antibiotics bind to PBPs, which are essential for bacterial cell wall synthesis. PBPs are members of a subgroup of enzymes called transpeptidases. Specifically, PBPs are DD-transpeptidases.

Panton–Valentine leukocidin (PVL) is a cytotoxin—one of the β-pore-forming toxins. The presence of PVL is associated with increased virulence of certain strains (isolates) of Staphylococcus aureus. It is present in the majority of community-associated methicillin-resistant Staphylococcus aureus (CA-MRSA) isolates studied and is the cause of necrotic lesions involving the skin or mucosa, including necrotic hemorrhagic pneumonia. PVL creates pores in the membranes of infected cells. PVL is produced from the genetic material of a bacteriophage that infects Staphylococcus aureus, making it more virulent.

Cefoxitin is a second-generation cephamycin antibiotic developed by Merck & Co., Inc. from Cephamycin C in the year following its discovery, 1972. It was synthesized in order to create an antibiotic with a broader spectrum. It is often grouped with the second-generation cephalosporins. Cefoxitin requires a prescription and as of 2010 is sold under the brand name Mefoxin by Bioniche Pharma, LLC. The generic version of cefoxitin is known as cefoxitin sodium.
Lysostaphin is a Staphylococcus simulans metalloendopeptidase. It can function as a bacteriocin (antimicrobial) against Staphylococcus aureus.

Beta-lactamases are a family of enzymes involved in bacterial resistance to beta-lactam antibiotics. In bacterial resistance to beta-lactam antibiotics, the bacteria have beta-lactamase which degrade the beta-lactam rings, rendering the antibiotic ineffective. However, with beta-lactamase inhibitors, these enzymes on the bacteria are inhibited, thus allowing the antibiotic to take effect. Strategies for combating this form of resistance have included the development of new beta-lactam antibiotics that are more resistant to cleavage and the development of the class of enzyme inhibitors called beta-lactamase inhibitors. Although β-lactamase inhibitors have little antibiotic activity of their own, they prevent bacterial degradation of beta-lactam antibiotics and thus extend the range of bacteria the drugs are effective against.

Ceftobiprole (Zevtera/Mabelio) is a fifth-generation cephalosporin for the treatment of hospital-acquired pneumonia and community-acquired pneumonia. It is marketed by Basilea Pharmaceutica in the United Kingdom, Germany, Switzerland and Austria under the trade name Zevtera, in France and Italy under the trade name Mabelio. Like other cephalosporins, ceftobiprole exerts its antibacterial activity by binding to important penicillin-binding proteins and inhibiting their transpeptidase activity which is essential for the synthesis of bacterial cell walls. Ceftobiprole has high affinity for penicillin-binding protein 2a of methicillin-resistant Staphylococcus aureus strains and retains its activity against strains that express divergent mecA gene homologues. Ceftobiprole also binds to penicillin-binding protein 2b in Streptococcus pneumoniae (penicillin-intermediate), to penicillin-binding protein 2x in Streptococcus pneumoniae (penicillin-resistant), and to penicillin-binding protein 5 in Enterococcus faecalis.
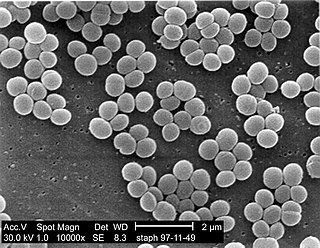
Staphylococcus is a genus of Gram-positive bacteria in the family Staphylococcaceae from the order Bacillales. Under the microscope, they appear spherical (cocci), and form in grape-like clusters. Staphylococcus species are facultative anaerobic organisms.
Cephalosporins are a broad class of bactericidal antibiotics that include the β-lactam ring and share a structural similarity and mechanism of action with other β-lactam antibiotics. The cephalosporins have the ability to kill bacteria by inhibiting essential steps in the bacterial cell wall synthesis which in the end results in osmotic lysis and death of the bacterial cell. Cephalosporins are widely used antibiotics because of their clinical efficiency and desirable safety profile.
SCCmec, or staphylococcal cassette chromosome mec, is a mobile genetic element of Staphylococcus bacterial species. This genetic sequence includes the mecA gene coding for resistance to the antibiotic methicillin and is the only known way for Staphylococcus strains to spread the gene in the wild by horizontal gene transfer. SCCmec is a 21 to 60 kb long genetic element that confers broad-spectrum β-lactam resistance to MRSA. Moreover, additional genetic elements like Tn554, pT181, and pUB110 can be found in SCCmec, which have the capability to render resistance to various non-β-lactam drugs.
Staphylococcus schleiferi is a Gram-positive, cocci-shaped bacterium of the family Staphylococcaceae. It is facultatively anaerobic, coagulase-variable, and can be readily cultured on blood agar where the bacterium tends to form opaque, non-pigmented colonies and beta (β) hemolysis. There exists two subspecies under the species S. schleiferi: Staphylococcus schleiferi subsp. schleiferi and Staphylococcus schleiferi subsp. coagulans.
Staphylococcus pseudintermedius is a gram positive coccus bacteria of the genus Staphylococcus found worldwide. It is primarily a pathogen for domestic animals, but has been known to affect humans as well. S. pseudintermedius is an opportunistic pathogen that secretes immune modulating virulence factors, has many adhesion factors, and the potential to create biofilms, all of which help to determine the pathogenicity of the bacterium. Diagnoses of Staphylococcus pseudintermedius have traditionally been made using cytology, plating, and biochemical tests. More recently, molecular technologies like MALDI-TOF, DNA hybridization and PCR have become preferred over biochemical tests for their more rapid and accurate identifications. This includes the identification and diagnosis of antibiotic resistant strains.
References
- ↑ Ubukata K, Nonoguchi R, Matsuhashi M, Konno M (May 1989). "Expression and inducibility in Staphylococcus aureus of the mecA gene, which encodes a methicillin-resistant S. aureus-specific penicillin-binding protein". Journal of Bacteriology. 171 (5): 2882–5. doi:10.1128/jb.171.5.2882-2885.1989. PMC 209980 . PMID 2708325.
- ↑ Deurenberg RH, Stobberingh EE (March 2009). "The molecular evolution of hospital- and community-associated methicillin-resistant Staphylococcus aureus". Current Molecular Medicine. 9 (2): 100–15. doi:10.2174/156652409787581637. PMID 19275621.
- ↑ Wielders CL, Fluit AC, Brisse S, Verhoef J, Schmitz FJ (November 2002). "mecA gene is widely disseminated in Staphylococcus aureus population". Journal of Clinical Microbiology. 40 (11): 3970–5. doi:10.1128/jcm.40.11.3970-3975.2002. PMC 139644 . PMID 12409360.
- ↑ Fogarty LR, Haack SK, Johnson HE, Brennan AK, Isaacs NM, Spencer C (2015). "Staphylococcus aureus and methicillin-resistant S. aureus (MRSA) at ambient freshwater beaches". Journal of Water and Health. 13 (3): 680–92. doi: 10.2166/wh.2014.278 . PMID 26322754.
- 1 2 Lowy FD (May 2003). "Antimicrobial resistance: the example of Staphylococcus aureus". The Journal of Clinical Investigation. 111 (9): 1265–73. doi:10.1172/JCI18535. PMC 154455 . PMID 12727914.
- ↑ Basset P, Feil EJ, Zanetti G, Blanc DS (2011). Tibayrenc M (ed.). Genetics and Evolution of Infectious Disease. London: Elsevier. pp. 669–688. doi:10.1016/B978-0-12-384890-1.00025-X. ISBN 9780123848901.
- 1 2 Katayama Y, Ito T, Hiramatsu K (June 2000). "A new class of genetic element, staphylococcus cassette chromosome mec, encodes methicillin resistance in Staphylococcus aureus". Antimicrobial Agents and Chemotherapy. 44 (6): 1549–55. doi:10.1128/aac.44.6.1549-1555.2000. PMC 89911 . PMID 10817707.
- ↑ "Global priority list of antibiotic-resistant bacteria to guide research, discovery, and development of new antibiotics". World Health Organization. Retrieved 2017-11-28.
- ↑ Ubukata K, Nakagami S, Nitta A, Yamane A, Kawakami S, Sugiura M, Konno M (July 1992). "Rapid detection of the mecA gene in methicillin-resistant staphylococci by enzymatic detection of polymerase chain reaction products". Journal of Clinical Microbiology. 30 (7): 1728–33. doi:10.1128/jcm.30.7.1728-1733.1992. PMC 265371 . PMID 1629327.
- ↑ Anand KB, Agrawal P, Kumar S, Kapila K (2009). "Comparison of cefoxitin disc diffusion test, oxacillin screen agar, and PCR for mecA gene for detection of MRSA". Indian Journal of Medical Microbiology. 27 (1): 27–9. doi: 10.1016/S0255-0857(21)01748-5 . hdl: 1807/53665 . PMID 19172055.
- ↑ Bignardi GE, Woodford N, Chapman A, Johnson AP, Speller DC (January 1996). "Detection of the mec-A gene and phenotypic detection of resistance in Staphylococcus aureus isolates with borderline or low-level methicillinresistance". The Journal of Antimicrobial Chemotherapy. 37 (1): 53–63. doi: 10.1093/jac/37.1.53 . PMID 8647774.
- ↑ Parvez MA, Shibata H, Nakano T, Niimi S, Fujii N, Arakaki N, Higuti T (August 2008). "No relationship exists between PBP 2a amounts expressed in different MRSA strains obtained clinically and their beta-lactam MIC values". The Journal of Medical Investigation. 55 (3–4): 246–53. doi: 10.2152/jmi.55.246 . PMID 18797139.
- ↑ Hanssen AM, Ericson Sollid JU (February 2006). "SCCmec in staphylococci: genes on the move". FEMS Immunology and Medical Microbiology. 46 (1): 8–20. doi: 10.1111/j.1574-695X.2005.00009.x . PMID 16420592.
- ↑ Hanssen AM, Sollid JU (May 2007). "Multiple staphylococcal cassette chromosomes and allelic variants of cassette chromosome recombinases in Staphylococcus aureus and coagulase-negative staphylococci from Norway". Antimicrobial Agents and Chemotherapy. 51 (5): 1671–7. doi:10.1128/AAC.00978-06. PMC 1855542 . PMID 17307983.
- ↑ Kuwahara-Arai K, Kondo N, Hori S, Tateda-Suzuki E, Hiramatsu K (December 1996). "Suppression of methicillin resistance in a mecA-containing pre-methicillin-resistant Staphylococcus aureus strain is caused by the mecI-mediated repression of PBP 2' production". Antimicrobial Agents and Chemotherapy. 40 (12): 2680–5. doi:10.1128/AAC.40.12.2680. PMC 163603 . PMID 9124822.
- ↑ Cano I, Alonso MC, Garcia-Rosado E, Saint-Jean SR, Castro D, Borrego JJ (March 2006). "Detection of lymphocystis disease virus (LCDV) in asymptomatic cultured gilt-head seabream (Sparus aurata, L.) using an immunoblot technique". Veterinary Microbiology. 113 (1–2): 137–41. doi:10.1016/j.vetmic.2005.10.024. PMID 16298500.
- ↑ Takeuchi F, Watanabe S, Baba T, Yuzawa H, Ito T, Morimoto Y, Kuroda M, Cui L, Takahashi M, Ankai A, Baba S, Fukui S, Lee JC, Hiramatsu K (November 2005). "Whole-genome sequencing of staphylococcus haemolyticus uncovers the extreme plasticity of its genome and the evolution of human-colonizing staphylococcal species". Journal of Bacteriology. 187 (21): 7292–308. doi:10.1128/JB.187.21.7292-7308.2005. PMC 1272970 . PMID 16237012.
- 1 2 Stapleton PD, Taylor PW (2002-02-15). "Methicillin resistance in Staphylococcus aureus: mechanisms and modulation". Science Progress. 85 (Pt 1): 57–72. doi:10.3184/003685002783238870. PMC 2065735 . PMID 11969119.
- ↑ Reed P, Veiga H, Jorge AM, Terrak M, Pinho MG (May 2011). "Monofunctional transglycosylases are not essential for Staphylococcus aureus cell wall synthesis". Journal of Bacteriology. 193 (10): 2549–56. doi:10.1128/JB.01474-10. PMC 3133172 . PMID 21441517.
- 1 2 Pinho MG, de Lencastre H, Tomasz A (September 2001). "An acquired and a native penicillin-binding protein cooperate in building the cell wall of drug-resistant staphylococci". Proceedings of the National Academy of Sciences of the United States of America. 98 (19): 10886–91. Bibcode:2001PNAS...9810886P. doi: 10.1073/pnas.191260798 . PMC 58569 . PMID 11517340.
- 1 2 Guignard B, Entenza JM, Moreillon P (October 2005). "Beta-lactams against methicillin-resistant Staphylococcus aureus". Current Opinion in Pharmacology. Anti-infectives/New technologies. 5 (5): 479–89. doi:10.1016/j.coph.2005.06.002. PMID 16095969.
- ↑ Huber J, Donald RG, Lee SH, Jarantow LW, Salvatore MJ, Meng X, Painter R, Onishi RH, Occi J, Dorso K, Young K, Park YW, Skwish S, Szymonifka MJ, Waddell TS, Miesel L, Phillips JW, Roemer T (August 2009). "Chemical genetic identification of peptidoglycan inhibitors potentiating carbapenem activity against methicillin-resistant Staphylococcus aureus". Chemistry & Biology. 16 (8): 837–48. doi: 10.1016/j.chembiol.2009.05.012 . PMID 19716474.
- ↑ Bernal P, Lemaire S, Pinho MG, Mobashery S, Hinds J, Taylor PW (July 2010). "Insertion of epicatechin gallate into the cytoplasmic membrane of methicillin-resistant Staphylococcus aureus disrupts penicillin-binding protein (PBP) 2a-mediated beta-lactam resistance by delocalizing PBP2". The Journal of Biological Chemistry. 285 (31): 24055–65. doi: 10.1074/jbc.M110.114793 . PMC 2911331 . PMID 20516078.
- ↑ Lee SH, Jarantow LW, Wang H, Sillaots S, Cheng H, Meredith TC, Thompson J, Roemer T (November 2011). "Antagonism of chemical genetic interaction networks resensitize MRSA to β-lactam antibiotics". Chemistry & Biology. 18 (11): 1379–89. doi: 10.1016/j.chembiol.2011.08.015 . PMID 22118672.
- ↑ Memmi G, Filipe SR, Pinho MG, Fu Z, Cheung A (November 2008). "Staphylococcus aureus PBP4 is essential for beta-lactam resistance in community-acquired methicillin-resistant strains". Antimicrobial Agents and Chemotherapy. 52 (11): 3955–66. doi:10.1128/AAC.00049-08. PMC 2573147 . PMID 18725435.
- ↑ Fuda C, Suvorov M, Shi Q, Hesek D, Lee M, Mobashery S (July 2007). "Shared functional attributes between the mecA gene product of Staphylococcus sciuri and penicillin-binding protein 2a of methicillin-resistant Staphylococcus aureus". Biochemistry. 46 (27): 8050–7. doi:10.1021/bi7004587. PMID 17567045.
- ↑ Severin A, Wu SW, Tabei K, Tomasz A (October 2005). "High-level (beta)-lactam resistance and cell wall synthesis catalyzed by the mecA homologue of Staphylococcus sciuri introduced into Staphylococcus aureus". Journal of Bacteriology. 187 (19): 6651–8. doi:10.1128/JB.187.19.6651-6658.2005. PMC 1251583 . PMID 16166526.
- ↑ Tsubakishita S, Kuwahara-Arai K, Sasaki T, Hiramatsu K (October 2010). "Origin and molecular evolution of the determinant of methicillin resistance in staphylococci". Antimicrobial Agents and Chemotherapy. 54 (10): 4352–9. doi:10.1128/AAC.00356-10. PMC 2944575 . PMID 20679504.