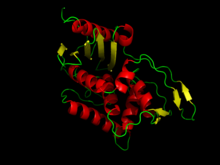
A beta-lactam (β-lactam) ring is a four-membered lactam. A lactam is a cyclic amide, and beta-lactams are named so because the nitrogen atom is attached to the β-carbon atom relative to the carbonyl. The simplest β-lactam possible is 2-azetidinone. β-lactams are significant structural units of medicines as manifested in many β-lactam antibiotics Up to 1970, most β-lactam research was concerned with the penicillin and cephalosporin groups, but since then, a wide variety of structures have been described.

Beta-lactamases (β-lactamases) are enzymes produced by bacteria that provide multi-resistance to beta-lactam antibiotics such as penicillins, cephalosporins, cephamycins, monobactams and carbapenems (ertapenem), although carbapenems are relatively resistant to beta-lactamase. Beta-lactamase provides antibiotic resistance by breaking the antibiotics' structure. These antibiotics all have a common element in their molecular structure: a four-atom ring known as a beta-lactam (β-lactam) ring. Through hydrolysis, the enzyme lactamase breaks the β-lactam ring open, deactivating the molecule's antibacterial properties.
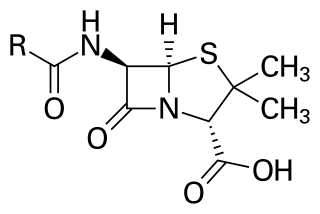
Penicillins are a group of β-lactam antibiotics originally obtained from Penicillium moulds, principally P. chrysogenum and P. rubens. Most penicillins in clinical use are synthesised by P. chrysogenum using deep tank fermentation and then purified. A number of natural penicillins have been discovered, but only two purified compounds are in clinical use: penicillin G and penicillin V. Penicillins were among the first medications to be effective against many bacterial infections caused by staphylococci and streptococci. They are still widely used today for different bacterial infections, though many types of bacteria have developed resistance following extensive use.
Peptidoglycan or murein is a unique large macromolecule, a polysaccharide, consisting of sugars and amino acids that forms a mesh-like peptidoglycan layer (sacculus) that surrounds the bacterial cytoplasmic membrane. The sugar component consists of alternating residues of β-(1,4) linked N-acetylglucosamine (NAG) and N-acetylmuramic acid (NAM). Attached to the N-acetylmuramic acid is an oligopeptide chain made of three to five amino acids. The peptide chain can be cross-linked to the peptide chain of another strand forming the 3D mesh-like layer. Peptidoglycan serves a structural role in the bacterial cell wall, giving structural strength, as well as counteracting the osmotic pressure of the cytoplasm. This repetitive linking results in a dense peptidoglycan layer which is critical for maintaining cell form and withstanding high osmotic pressures, and it is regularly replaced by peptidoglycan production. Peptidoglycan hydrolysis and synthesis are two processes that must occur in order for cells to grow and multiply, a technique carried out in three stages: clipping of current material, insertion of new material, and re-crosslinking of existing material to new material.
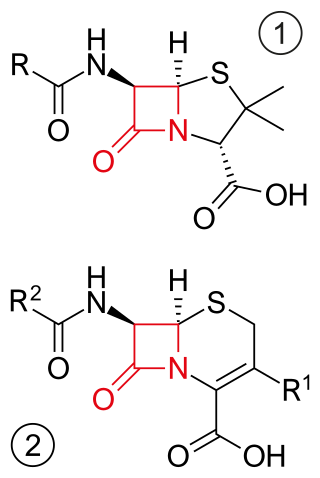
β-lactam antibiotics are antibiotics that contain a beta-lactam ring in their chemical structure. This includes penicillin derivatives (penams), cephalosporins and cephamycins (cephems), monobactams, carbapenems and carbacephems. Most β-lactam antibiotics work by inhibiting cell wall biosynthesis in the bacterial organism and are the most widely used group of antibiotics. Until 2003, when measured by sales, more than half of all commercially available antibiotics in use were β-lactam compounds. The first β-lactam antibiotic discovered, penicillin, was isolated from a strain of Penicillium rubens.

Methicillin (USAN), also known as meticillin (INN), is a narrow-spectrum β-lactam antibiotic of the penicillin class.

Clavulanic acid is a β-lactam drug that functions as a mechanism-based β-lactamase inhibitor. While not effective by itself as an antibiotic, when combined with penicillin-group antibiotics, it can overcome antibiotic resistance in bacteria that secrete β-lactamase, which otherwise inactivates most penicillins.

The acyl carrier protein (ACP) is a cofactor of both fatty acid and polyketide biosynthesis machinery. It is one of the most abundant proteins in cells of E. coli. In both cases, the growing chain is bound to the ACP via a thioester derived from the distal thiol of a 4'-phosphopantetheine moiety.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.
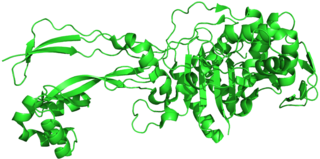
Penicillin-binding proteins (PBPs) are a group of proteins that are characterized by their affinity for and binding of penicillin. They are a normal constituent of many bacteria; the name just reflects the way by which the protein was discovered. All β-lactam antibiotics bind to PBPs, which are essential for bacterial cell wall synthesis. PBPs are members of a subgroup of enzymes called transpeptidases. Specifically, PBPs are DD-transpeptidases.
A carboxypeptidase is a protease enzyme that hydrolyzes (cleaves) a peptide bond at the carboxy-terminal (C-terminal) end of a protein or peptide. This is in contrast to an aminopeptidases, which cleave peptide bonds at the N-terminus of proteins. Humans, animals, bacteria and plants contain several types of carboxypeptidases that have diverse functions ranging from catabolism to protein maturation. At least two mechanisms have been discussed.

Beta-lactamases are a family of enzymes involved in bacterial resistance to beta-lactam antibiotics. In bacterial resistance to beta-lactam antibiotics, the bacteria have beta-lactamase which degrade the beta-lactam rings, rendering the antibiotic ineffective. However, with beta-lactamase inhibitors, these enzymes on the bacteria are inhibited, thus allowing the antibiotic to take effect. Strategies for combating this form of resistance have included the development of new beta-lactam antibiotics that are more resistant to cleavage and the development of the class of enzyme inhibitors called beta-lactamase inhibitors. Although β-lactamase inhibitors have little antibiotic activity of their own, they prevent bacterial degradation of beta-lactam antibiotics and thus extend the range of bacteria the drugs are effective against.
The discovery of an orally inactive peptide from snake venom established the important role of angiotensin converting enzyme (ACE) inhibitors in regulating blood pressure. This led to the development of captopril, the first ACE inhibitor. When the adverse effects of captopril became apparent new derivates were designed. Then after the discovery of two active sites of ACE: N-domain and C-domain, the development of domain-specific ACE inhibitors began.
Jack Leonard Strominger is the Higgins Professor of Biochemistry at Harvard University, specializing in the structure and function of human histocompatibility proteins and their role in disease. He won the Albert Lasker Award for Basic Medical Research in 1995.

Peptidoglycan binding domains have a general peptidoglycan binding function and a common core structure consisting of a closed, three-helical bundle with a left-handed twist. It is found at the N or C terminus of a variety of enzymes involved in bacterial cell wall degradation. Examples are:
Cephalosporins are a broad class of bactericidal antibiotics that include the β-lactam ring and share a structural similarity and mechanism of action with other β-lactam antibiotics. The cephalosporins have the ability to kill bacteria by inhibiting essential steps in the bacterial cell wall synthesis which in the end results in osmotic lysis and death of the bacterial cell. Cephalosporins are widely used antibiotics because of their clinical efficiency and desirable safety profile.
In molecular biology, VanY are protein domains found in enzymes named metallopeptidases. They are vital to bacterial cell wall synthesis and antibiotic resistance.
Muramoylpentapeptide carboxypeptidase is an enzyme. This enzyme catalyses the following chemical reaction.
Muramoyltetrapeptide carboxypeptidase is an enzyme. This enzyme catalyses the following chemical reaction
Zinc D-Ala-D-Ala carboxypeptidase (EC 3.4.17.14, Zn2+ G peptidase, D-alanyl-D-alanine hydrolase, D-alanyl-D-alanine-cleaving carboxypeptidase, DD-carboxypeptidase, G enzyme, DD-carboxypeptidase-transpeptidase) is an enzyme. This enzyme catalyses the following chemical reaction