Related Research Articles
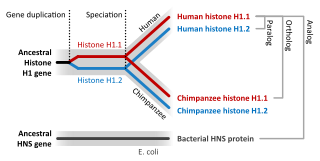
Sequence homology is the biological homology between DNA, RNA, or protein sequences, defined in terms of shared ancestry in the evolutionary history of life. Two segments of DNA can have shared ancestry because of three phenomena: either a speciation event (orthologs), or a duplication event (paralogs), or else a horizontal gene transfer event (xenologs).
The Gene Ontology (GO) is a major bioinformatics initiative to unify the representation of gene and gene product attributes across all species. More specifically, the project aims to: 1) maintain and develop its controlled vocabulary of gene and gene product attributes; 2) annotate genes and gene products, and assimilate and disseminate annotation data; and 3) provide tools for easy access to all aspects of the data provided by the project, and to enable functional interpretation of experimental data using the GO, for example via enrichment analysis. GO is part of a larger classification effort, the Open Biomedical Ontologies, being one of the Initial Candidate Members of the OBO Foundry.

Ensembl genome database project is a scientific project at the European Bioinformatics Institute, which provides a centralized resource for geneticists, molecular biologists and other researchers studying the genomes of our own species and other vertebrates and model organisms. Ensembl is one of several well known genome browsers for the retrieval of genomic information.
The Rat Genome Database (RGD) is a database of rat genomics, genetics, physiology and functional data, as well as data for comparative genomics between rat, human and mouse. RGD is responsible for attaching biological information to the rat genome via structured vocabulary, or ontology, annotations assigned to genes and quantitative trait loci (QTL), and for consolidating rat strain data and making it available to the research community. They are also developing a suite of tools for mining and analyzing genomic, physiologic and functional data for the rat, and comparative data for rat, mouse, human, and five other species.

KEGG is a collection of databases dealing with genomes, biological pathways, diseases, drugs, and chemical substances. KEGG is utilized for bioinformatics research and education, including data analysis in genomics, metagenomics, metabolomics and other omics studies, modeling and simulation in systems biology, and translational research in drug development.

The Biological General Repository for Interaction Datasets (BioGRID) is a curated biological database of protein-protein interactions, genetic interactions, chemical interactions, and post-translational modifications created in 2003 (originally referred to as simply the General Repository for Interaction Datasets by Mike Tyers, Bobby-Joe Breitkreutz, and Chris Stark at the Lunenfeld-Tanenbaum Research Institute at Mount Sinai Hospital. It strives to provide a comprehensive curated resource for all major model organism species while attempting to remove redundancy to create a single mapping of data. Users of The BioGRID can search for their protein, chemical or publication of interest and retrieve annotation, as well as curated data as reported, by the primary literature and compiled by in house large-scale curation efforts. The BioGRID is hosted in Toronto, Ontario, Canada and Dallas, Texas, United States and is partnered with the Saccharomyces Genome Database, FlyBase, WormBase, PomBase, and the Alliance of Genome Resources. The BioGRID is funded by the NIH and CIHR. BioGRID is an observer member of the International Molecular Exchange Consortium.
The Saccharomyces Genome Database (SGD) is a scientific database of the molecular biology and genetics of the yeast Saccharomyces cerevisiae, which is commonly known as baker's or budding yeast. Further information is located at the Yeastract curated repository.

The Generic Model Organism Database (GMOD) project provides biological research communities with a toolkit of open-source software components for visualizing, annotating, managing, and storing biological data. The GMOD project is funded by the United States National Institutes of Health, National Science Foundation and the USDA Agricultural Research Service.
FlyBase is an online bioinformatics database and the primary repository of genetic and molecular data for the insect family Drosophilidae. For the most extensively studied species and model organism, Drosophila melanogaster, a wide range of data are presented in different formats.
Mouse Genome Informatics (MGI) is a free, online database and bioinformatics resource hosted by The Jackson Laboratory, with funding by the National Human Genome Research Institute (NHGRI), the National Cancer Institute (NCI), and the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD). MGI provides access to data on the genetics, genomics and biology of the laboratory mouse to facilitate the study of human health and disease. The database integrates multiple projects, with the two largest contributions coming from the Mouse Genome Database and Mouse Gene Expression Database (GXD). As of 2018, MGI contains data curated from over 230,000 publications.

MicrobesOnline is a publicly and freely accessible website that hosts multiple comparative genomic tools for comparing microbial species at the genomic, transcriptomic and functional levels. MicrobesOnline was developed by the Virtual Institute for Microbial Stress and Survival, which is based at the Lawrence Berkeley National Laboratory in Berkeley, California. The site was launched in 2005, with regular updates until 2011.
The Reference Sequence (RefSeq) database is an open access, annotated and curated collection of publicly available nucleotide sequences and their protein products. RefSeq was introduced in 2000. This database is built by National Center for Biotechnology Information (NCBI), and, unlike GenBank, provides only a single record for each natural biological molecule for major organisms ranging from viruses to bacteria to eukaryotes.
SUPERFAMILY is a database and search platform of structural and functional annotation for all proteins and genomes. It classifies amino acid sequences into known structural domains, especially into SCOP superfamilies. Domains are functional, structural, and evolutionary units that form proteins. Domains of common Ancestry are grouped into superfamilies. The domains and domain superfamilies are defined and described in SCOP. Superfamilies are groups of proteins which have structural evidence to support a common evolutionary ancestor but may not have detectable sequence homology.
Protein function prediction methods are techniques that bioinformatics researchers use to assign biological or biochemical roles to proteins. These proteins are usually ones that are poorly studied or predicted based on genomic sequence data. These predictions are often driven by data-intensive computational procedures. Information may come from nucleic acid sequence homology, gene expression profiles, protein domain structures, text mining of publications, phylogenetic profiles, phenotypic profiles, and protein-protein interaction. Protein function is a broad term: the roles of proteins range from catalysis of biochemical reactions to transport to signal transduction, and a single protein may play a role in multiple processes or cellular pathways.

In molecular biology and genetics, DNA annotation or genome annotation is the process of describing the structure and function of the components of a genome, by analyzing and interpreting them in order to extract their biological significance and understand the biological processes in which they participate. Among other things, it identifies the locations of genes and all the coding regions in a genome and determines what those genes do.
In bioinformatics, the PANTHER classification system is a large curated biological database of gene/protein families and their functionally related subfamilies that can be used to classify and identify the function of gene products. PANTHER is part of the Gene Ontology Reference Genome Project designed to classify proteins and their genes for high-throughput analysis.
PomBase is a model organism database that provides online access to the fission yeast Schizosaccharomyces pombe genome sequence and annotated features, together with a wide range of manually curated functional gene-specific data. The PomBase website was redeveloped in 2016 to provide users with a more fully integrated, better-performing service.
Judith Anne Blake is a computational biologist at the Jackson Laboratory and Professor of Mammalian Genetics.
Biocuration is the field of life sciences dedicated to organizing biomedical data, information and knowledge into structured formats, such as spreadsheets, tables and knowledge graphs. The biocuration of biomedical knowledge is made possible by the cooperative work of biocurators, software developers and bioinformaticians and is at the base of the work of biological databases.
References
- 1 2 3 Oliver SG, Lock A, Harris MA, Nurse P, Wood V (June 2016). "Model organism databases: essential resources that need the support of both funders and users". BMC Biology. 14 (1): 49. doi: 10.1186/s12915-016-0276-z . PMC 4918006 . PMID 27334346.
- ↑ Bond M, Holthaus SM, Tammen I, Tear G, Russell C (November 2013). "Use of model organisms for the study of neuronal ceroid lipofuscinosis". Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease. 1832 (11): 1842–65. doi: 10.1016/j.bbadis.2013.01.009 . PMID 23338040.
- ↑ Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. (May 2000). "Gene ontology: tool for the unification of biology. The Gene Ontology Consortium". Nature Genetics. 25 (1): 25–9. doi:10.1038/75556. PMC 3037419 . PMID 10802651.
- ↑ Gene Ontology Consortium (January 2015). "Gene Ontology Consortium: going forward". Nucleic Acids Research. 43 (Database issue): D1049-56. doi:10.1093/nar/gku1179. PMC 4383973 . PMID 25428369.
- ↑ O'Connor BD, Day A, Cain S, Arnaiz O, Sperling L, Stein LD (2008). "GMODWeb: a web framework for the Generic Model Organism Database". Genome Biology. 9 (6): R102. doi: 10.1186/gb-2008-9-6-r102 . PMC 2481422 . PMID 18570664.
- ↑ Cherry JM, Hong EL, Amundsen C, Balakrishnan R, Binkley G, Chan ET, et al. (January 2012). "Saccharomyces Genome Database: the genomics resource of budding yeast". Nucleic Acids Research. 40 (Database issue): D700-5. doi:10.1093/nar/gkr1029. PMC 3245034 . PMID 22110037.
- ↑ Lock A, Rutherford K, Harris MA, Hayles J, Oliver SG, Bähler J, Wood V (January 2019). "PomBase 2018: user-driven reimplementation of the fission yeast database provides rapid and intuitive access to diverse, interconnected information". Nucleic Acids Research. 47 (D1): D821–D827. doi:10.1093/nar/gky961. PMC 6324063 . PMID 30321395.
- ↑ Wood V, Harris MA, McDowall MD, Rutherford K, Vaughan BW, Staines DM, et al. (January 2012). "PomBase: a comprehensive online resource for fission yeast". Nucleic Acids Research. 40 (Database issue): D695-9. doi:10.1093/nar/gkr853. PMC 3245111 . PMID 22039153.
- ↑ McDowall MD, Harris MA, Lock A, Rutherford K, Staines DM, Bähler J, et al. (January 2015). "PomBase 2015: updates to the fission yeast database". Nucleic Acids Research. 43 (Database issue): D656-61. doi:10.1093/nar/gku1040. PMC 4383888 . PMID 25361970.
- ↑ Lock A, Rutherford K, Harris MA, Wood V (2018). "PomBase: The Scientific Resource for Fission Yeast". Eukaryotic Genomic Databases. Methods in Molecular Biology. Vol. 1757. pp. 49–68. doi:10.1007/978-1-4939-7737-6_4. ISBN 978-1-4939-7736-9. PMC 6440643 . PMID 29761456.
- ↑ Karimi K, Fortriede JD, Lotay VS, Burns KA, Wang DZ, Fisher ME, et al. (January 2018). "Xenbase: a genomic, epigenomic and transcriptomic model organism database". Nucleic Acids Research. 46 (D1): D861–D868. doi:10.1093/nar/gkx936. PMC 5753396 . PMID 29059324.
- ↑ James-Zorn C, Ponferrada VG, Fisher ME, Burns KA, Fortriede JD, Segerdell E, Karimi K, Lotay VS, Wang DZ, Chu S, Pells TJ, Wang Y, Vize PD, Zorn AM (May 2018). "Navigating Xenbase: An Integrated Xenopus Genomics and Gene Expression Database". Eukaryotic Genomic Databases. Methods in Molecular Biology. Vol. 1757. pp. 251–305. doi:10.1007/978-1-4939-7737-6_10. ISBN 978-1-4939-7736-9. PMC 6853059 . PMID 29761462.
{{cite book}}:|journal=ignored (help) - ↑ Telmer CA, Karimi K, Chess MM, Agalakov S, Arshinoff BI, Lotay V, Wang DZ, Chu S, Pells TJ, Vize PD, Hinman VF, Ettensohn CA, Echinobase: a resource to support the echinoderm research community, Genetics, 2024, doi:10.1093/genetics/iyae002
- ↑ Attrill H, Falls K, Goodman JL, Millburn GH, Antonazzo G, Rey AJ, Marygold SJ (January 2016). "FlyBase: establishing a Gene Group resource for Drosophila melanogaster". Nucleic Acids Research. 44 (D1): D786-92. doi:10.1093/nar/gkv1046. PMC 4702782 . PMID 26467478.
- ↑ Elsik CG, Tayal A, Unni DR, Burns GW, Hagen DE (2018). "Hymenoptera Genome Database: Using HymenopteraMine to Enhance Genomic Studies of Hymenopteran Insects". In Kollmar M (ed.). Eukaryotic Genomic Databases. Methods in Molecular Biology. Vol. 1757. New York, NY: Springer New York. pp. 513–556. doi:10.1007/978-1-4939-7737-6_17. ISBN 978-1-4939-7736-9. PMID 29761469.
- ↑ Eppig JT, Blake JA, Bult CJ, Kadin JA, Richardson JE (January 2015). "The Mouse Genome Database (MGD): facilitating mouse as a model for human biology and disease". Nucleic Acids Research. 43 (Database issue): D726-36. doi:10.1093/nar/gku967. PMC 4384027 . PMID 25348401.
- ↑ Harris TW, Baran J, Bieri T, Cabunoc A, Chan J, Chen WJ, et al. (January 2014). "WormBase 2014: new views of curated biology". Nucleic Acids Research. 42 (Database issue): D789-93. doi:10.1093/nar/gkt1063. PMC 3965043 . PMID 24194605.
- ↑ Shimoyama M, De Pons J, Hayman GT, Laulederkind SJ, Liu W, Nigam R, et al. (January 2015). "The Rat Genome Database 2015: genomic, phenotypic and environmental variations and disease". Nucleic Acids Research. 43 (Database issue): D743-50. doi:10.1093/nar/gku1026. PMC 4383884 . PMID 25355511.
- ↑ Kreppel L, Fey P, Gaudet P, Just E, Kibbe WA, Chisholm RL, Kimmel AR (January 2004). "dictyBase: a new Dictyostelium discoideum genome database". Nucleic Acids Research. 32 (Database issue): D332-3. doi:10.1093/nar/gkh138. PMC 308872 . PMID 14681427.
- ↑ Lamesch P, Berardini TZ, Li D, Swarbreck D, Wilks C, Sasidharan R, et al. (January 2012). "The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools". Nucleic Acids Research. 40 (Database issue): D1202-10. doi:10.1093/nar/gkr1090. PMC 3245047 . PMID 22140109.
- ↑ Lawrence CJ, Dong Q, Polacco ML, Seigfried TE, Brendel V (January 2004). "MaizeGDB, the community database for maize genetics and genomics". Nucleic Acids Research. 32 (Database issue): D393-7. doi:10.1093/nar/gkh011. PMC 308746 . PMID 14681441.
- ↑ Andorf CM, Cannon EK, Portwood JL, Gardiner JM, Harper LC, Schaeffer ML, et al. (January 2016). "MaizeGDB update: new tools, data and interface for the maize model organism database". Nucleic Acids Research. 44 (D1): D1195-201. doi:10.1093/nar/gkv1007. PMC 4702771 . PMID 26432828.
- ↑ Grant D, Nelson RT, Cannon SB, Shoemaker RC (January 2010). "SoyBase, the USDA-ARS soybean genetics and genomics database". Nucleic Acids Research. 38 (Database issue): D843-6. doi:10.1093/nar/gkp798. PMC 2808871 . PMID 20008513.
- ↑ Howe DG, Bradford YM, Conlin T, Eagle AE, Fashena D, Frazer K, et al. (January 2013). "ZFIN, the Zebrafish Model Organism Database: increased support for mutants and transgenics". Nucleic Acids Research. 41 (Database issue): D854-60. doi:10.1093/nar/gks938. PMC 3531097 . PMID 23074187.
- ↑ Inglis DO, Arnaud MB, Binkley J, Shah P, Skrzypek MS, Wymore F, et al. (January 2012). "The Candida genome database incorporates multiple Candida species: multispecies search and analysis tools with curated gene and protein information for Candida albicans and Candida glabrata". Nucleic Acids Research. 40 (Database issue): D667-74. doi:10.1093/nar/gkr945. PMC 3245171 . PMID 22064862.
- ↑ Keseler IM, Mackie A, Peralta-Gil M, Santos-Zavaleta A, Gama-Castro S, Bonavides-Martínez C, et al. (January 2013). "EcoCyc: fusing model organism databases with systems biology". Nucleic Acids Research. 41 (Database issue): D605-12. doi:10.1093/nar/gks1027. PMC 3531154 . PMID 23143106.
- ↑ Zhu B, Stülke J (January 2018). "SubtiWiki in 2018: from genes and proteins to functional network annotation of the model organism Bacillus subtilis". Nucleic Acids Research. 46 (D1): D743–D748. doi:10.1093/nar/gkx908. PMC 5753275 . PMID 29788229.