A metalloproteinase, or metalloprotease, is any protease enzyme whose catalytic mechanism involves a metal. An example is ADAM12 which plays a significant role in the fusion of muscle cells during embryo development, in a process known as myogenesis.
The cystatins are a family of cysteine protease inhibitors which share a sequence homology and a common tertiary structure of an alpha helix lying on top of an anti-parallel beta sheet. The family is subdivided as described below.

ADAMs are a family of single-pass transmembrane and secreted metalloendopeptidases. All ADAMs are characterized by a particular domain organization featuring a pro-domain, a metalloprotease, a disintegrin, a cysteine-rich, an epidermal-growth factor like and a transmembrane domain, as well as a C-terminal cytoplasmic tail. Nonetheless, not all human ADAMs have a functional protease domain, which indicates that their biological function mainly depends on protein–protein interactions. Those ADAMs which are active proteases are classified as sheddases because they cut off or shed extracellular portions of transmembrane proteins. For example, ADAM10 can cut off part of the HER2 receptor, thereby activating it. ADAM genes are found in animals, choanoflagellates, fungi and some groups of green algae. Most green algae and all land plants likely lost ADAM proteins.
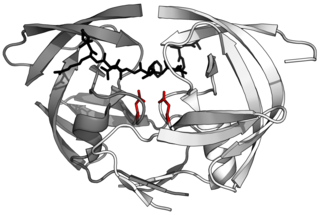
Aspartic proteases are a catalytic type of protease enzymes that use an activated water molecule bound to one or more aspartate residues for catalysis of their peptide substrates. In general, they have two highly conserved aspartates in the active site and are optimally active at acidic pH. Nearly all known aspartyl proteases are inhibited by pepstatin.
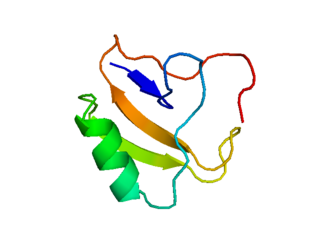
Scorpion toxins are proteins found in the venom of scorpions. Their toxic effect may be mammal- or insect-specific and acts by binding with varying degrees of specificity to members of the Voltage-gated ion channel superfamily; specifically, voltage-gated sodium channels, voltage-gated potassium channels, and Transient Receptor Potential (TRP) channels. The result of this action is to activate or inhibit the action of these channels in the nervous and cardiac organ systems. For instance, α-scorpion toxins MeuNaTxα-12 and MeuNaTxα-13 from Mesobuthus eupeus are neurotoxins that target voltage-gated Na+ channels (Navs), inhibiting fast inactivation. In vivo assays of MeuNaTxα-12 and MeuNaTxα-13 effects on mammalian and insect Navs show differential potency. These recombinants exhibit their preferential affinity for mammalian and insect Na+ channels at the α-like toxins' active site, site 3, in order to inactivate the cell membrane depolarization faster[6]. The varying sensitivity of different Navs to MeuNaTxα-12 and MeuNaTxα-13 may be dependent on the substitution of a conserved Valine residue for a Phenylalanine residue at position 1630 of the LD4:S3-S4 subunit or due to various changes in residues in the LD4:S5-S6 subunit of the Navs. Ultimately, these actions can serve the purpose of warding off predators by causing pain or to subdue predators.
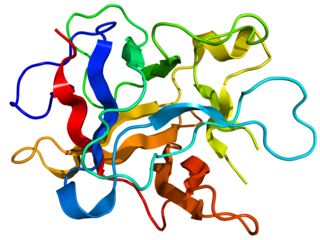
Kunitz soybean trypsin inhibitor is a type of protein contained in legume seeds which functions as a protease inhibitor. Kunitz-type Soybean Trypsin Inhibitors are usually specific for either trypsin or chymotrypsin. They are thought to protect seeds against consumption by animal predators.
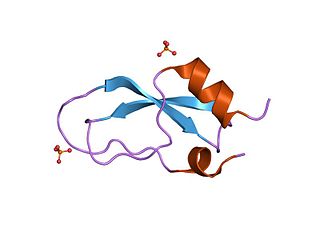
Kunitz domains are the active domains of proteins that inhibit the function of protein degrading enzymes or, more specifically, domains of Kunitz-type are protease inhibitors. They are relatively small with a length of about 50 to 60 amino acids and a molecular weight of 6 kDa. Examples of Kunitz-type protease inhibitors are aprotinin, Alzheimer's amyloid precursor protein (APP), and tissue factor pathway inhibitor (TFPI). Kunitz STI protease inhibitor, the trypsin inhibitor initially studied by Moses Kunitz, was extracted from soybeans.

In molecular biology, the CHAP domain is a region between 110 and 140 amino acids that is found in proteins from bacteria, bacteriophages, archaea and eukaryotes of the family Trypanosomidae. The domain is named after the acronym cysteine, histidine-dependent amidohydrolases/peptidases. Many of these proteins are uncharacterised, but it has been proposed that they may function mainly in peptidoglycan hydrolysis. The CHAP domain is found in a wide range of protein architectures; it is commonly associated with bacterial type SH3 domains and with several families of amidase domains. It has been suggested that CHAP domain containing proteins utilise a catalytic cysteine residue in a nucleophilic-attack mechanism.
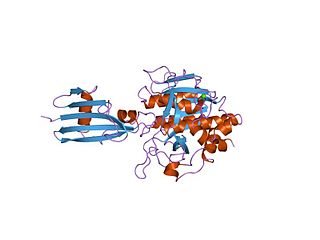
In molecular biology the protein SSI is a Subtilisin inhibitor-like which stands for Streptomyces subtilisin inhibitor. This is a protease inhibitor. These are often synthesised as part of a larger precursor protein, either as a prepropeptide. The function of this protein domain is to prevent access of the substrate to the active site. It is found only in bacteria.
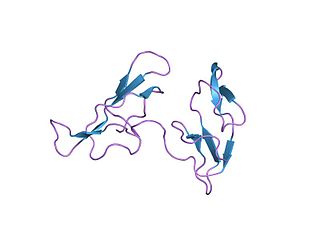
In molecular biology, the Bowman–Birk protease inhibitor family of proteins consists of eukaryotic proteinase inhibitors, belonging to MEROPS inhibitor family I12, clan IF. They mainly inhibit serine peptidases of the S1 family, but also inhibit S3 peptidases.
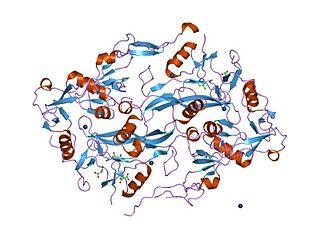
The Kazal domain is an evolutionary conserved protein domain usually indicative of serine protease inhibitors. However, kazal-like domains are also seen in the extracellular part of agrins, which are not known to be protease inhibitors.
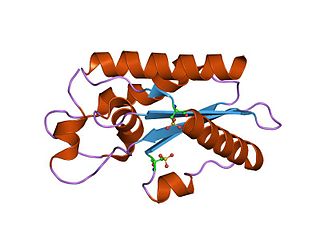
The CAP superfamily is a large superfamily of secreted proteins that are produced by a wide range of organisms, including prokaryotes and non-vertebrate eukaryotes.

In molecular biology, the carboxypeptidase A inhibitor family is a family of proteins which is represented by the well-characterised metallocarboxypeptidase A inhibitor (MCPI) from potatoes, which belongs to the MEROPS inhibitor family I37, clan IE. It inhibits metallopeptidases belonging to MEROPS peptidase family M14, carboxypeptidase A. In Russet Burbank potatoes, it is a mixture of approximately equal amounts of two polypeptide chains containing 38 or 39 amino acid residues. The chains differ in their amino terminal sequence only and are resistant to fragmentation by proteases. The structure of the complex between bovine carboxypeptidase A and the 39-amino-acid carboxypeptidase A inhibitor from potatoes has been determined at 2.5-Angstrom resolution.

In molecular biology, the trappin protein transglutaminase binding domain or cementoin is a protein domain found at the N-terminus of Whey Acidic Protein (WAP) domain-containing protease inhibitors such as trappin-2. This N-terminal domain enables it to become cross-linked to extracellular matrix proteins by transglutaminase. This domain contains several repeated motifs with the consensus sequence Gly-Gln-Asp-Pro-Val-Lys, and these together can anchor the whole molecule to extracellular matrix proteins, such as laminin, fibronectin, beta-crystallin, collagen IV, fibrinogen, and elastin, by transglutaminase-catalysed cross-links. The whole domain is rich in glutamine and lysine, thus allowing transglutaminase(s) to catalyse the formation of an intermolecular epsilon-(gamma-glutamyl)lysine isopeptide bond.
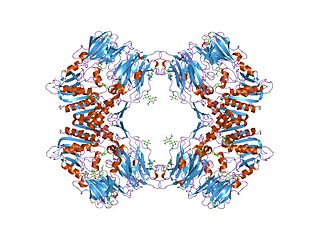

In molecular biology, the dipeptidyl-peptidase IV family is a family of serine peptidases which belong to MEROPS peptidase family S9, subfamily S9B. The protein fold of the peptidase domain for members of this family resembles that of serine carboxypeptidase D, the type example of clan SC. The type example of this family is Dipeptidyl peptidase-4.

In molecular biology, ecotin is a protease inhibitor which belongs to MEROPS inhibitor family I11, clan IN. Ecotins are dimeric periplasmic proteins from Escherichia coli and related Gram-negative bacteria that have been shown to be potent inhibitors of many trypsin-fold serine proteases of widely varying substrate specificity, which belong to MEROPS peptidase family S1. Phylogenetic analysis suggested that ecotin has an exogenous target, possibly neutrophil elastase. Ecotin from E. coli, Yersinia pestis, and Pseudomonas aeruginosa, all species that encounter the mammalian immune system, inhibit neutrophil elastase strongly while ecotin from the plant pathogen Pantoea citrea inhibits neutrophil elastase 1000-fold less potently. Ecotins all potently inhibit pancreatic digestive peptidases trypsin and chymotrypsin, while showing more variable inhibition of the blood peptidases Factor Xa, thrombin, and urokinase-type plasminogen activator.

In molecular biology, haemadin is an anticoagulant peptide synthesised by the Indian leech, Haemadipsa sylvestris. It adopts a secondary structure consisting of five short beta-strands (beta1-beta5), which are arranged in two antiparallel distorted sheets formed by strands beta1-beta4-beta5 and beta2-beta3 facing each other. This beta-sandwich is stabilised by six enclosed cysteines arranged in a [1-2, 3-5, 4-6] disulfide pairing resulting in a disulfide-rich hydrophobic core that is largely inaccessible to bulk solvent. The close proximity of disulfide bonds [3-5] and [4-6] organises haemadin into four distinct loops. The N-terminal segment of this domain binds to the active site of thrombin, inhibiting it.

In molecular biology, the cyanobacterial clock proteins are the main circadian regulator in cyanobacteria. The cyanobacterial clock proteins comprise three proteins: KaiA, KaiB and KaiC. The kaiABC complex may act as a promoter-nonspecific transcription regulator that represses transcription, possibly by acting on the state of chromosome compaction. This complex is expressed from a KaiABC operon.

The 3C-like protease (3CLpro) or main protease (Mpro), formally known as C30 endopeptidase or 3-chymotrypsin-like protease, is the main protease found in coronaviruses. It cleaves the coronavirus polyprotein at eleven conserved sites. It is a cysteine protease and a member of the PA clan of proteases. It has a cysteine-histidine catalytic dyad at its active site and cleaves a Gln–(Ser/Ala/Gly) peptide bond.

Pacifastin is a family of serine proteinase inhibitors found in arthropods. Pacifastin inhibits the serine peptidases trypsin and chymotrypsin.