Related Research Articles

A polyhistidine-tag, best known by the trademarked name His-tag, is an amino acid motif in proteins that typically consists of at least six histidine (His) residues, often at the N- or C-terminus of the protein. It is also known as a hexa histidine-tag, 6xHis-tag, or His6 tag. The tag was invented by Roche, although the use of histidines and its vectors are distributed by Qiagen. Various purification kits for histidine-tagged proteins are commercially available from multiple companies.
An epitope, also known as antigenic determinant, is the part of an antigen that is recognized by the immune system, specifically by antibodies, B cells, or T cells. The part of an antibody that binds to the epitope is called a paratope. Although epitopes are usually non-self proteins, sequences derived from the host that can be recognized are also epitopes.
Protein purification is a series of processes intended to isolate one or a few proteins from a complex mixture, usually cells, tissues or whole organisms. Protein purification is vital for the specification of the function, structure and interactions of the protein of interest. The purification process may separate the protein and non-protein parts of the mixture, and finally separate the desired protein from all other proteins. Ideally, to study a protein of interest, it must be separated from other components of the cell so that contaminants will not interfere in the examination of the protein of interest's structure and function. Separation of one protein from all others is typically the most laborious aspect of protein purification. Separation steps usually exploit differences in protein size, physico-chemical properties, binding affinity and biological activity. The pure result may be termed protein isolate.
In biochemistry, biotinylation is the process of covalently attaching biotin to a protein, nucleic acid or other molecule. Biotinylation is rapid, specific and is unlikely to disturb the natural function of the molecule due to the small size of biotin. Biotin binds to streptavidin and avidin with an extremely high affinity, fast on-rate, and high specificity, and these interactions are exploited in many areas of biotechnology to isolate biotinylated molecules of interest. Biotin-binding to streptavidin and avidin is resistant to extremes of heat, pH and proteolysis, making capture of biotinylated molecules possible in a wide variety of environments. Also, multiple biotin molecules can be conjugated to a protein of interest, which allows binding of multiple streptavidin, avidin or neutravidin protein molecules and increases the sensitivity of detection of the protein of interest. There is a large number of biotinylation reagents available that exploit the wide range of possible labelling methods. Due to the strong affinity between biotin and streptavidin, the purification of biotinylated proteins has been a widely used approach to identify protein-protein interactions and post-translational events such as ubiquitylation in molecular biology.
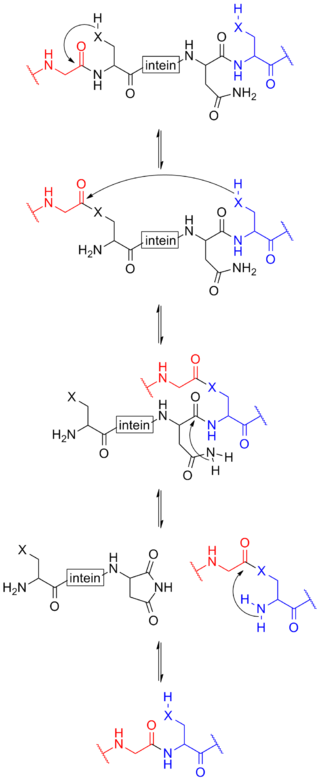
Protein splicing is an intramolecular reaction of a particular protein in which an internal protein segment is removed from a precursor protein with a ligation of C-terminal and N-terminal external proteins on both sides. The splicing junction of the precursor protein is mainly a cysteine or a serine, which are amino acids containing a nucleophilic side chain. The protein splicing reactions which are known now do not require exogenous cofactors or energy sources such as adenosine triphosphate (ATP) or guanosine triphosphate (GTP). Normally, splicing is associated only with pre-mRNA splicing. This precursor protein contains three segments—an N-extein followed by the intein followed by a C-extein. After splicing has taken place, the resulting protein contains the N-extein linked to the C-extein; this splicing product is also termed an extein.
Affinity chromatography is a method of separating a biomolecule from a mixture, based on a highly specific macromolecular binding interaction between the biomolecule and another substance. The specific type of binding interaction depends on the biomolecule of interest; antigen and antibody, enzyme and substrate, receptor and ligand, or protein and nucleic acid binding interactions are frequently exploited for isolation of various biomolecules. Affinity chromatography is useful for its high selectivity and resolution of separation, compared to other chromatographic methods.

Streptavidin is a 52 kDa protein (tetramer) purified from the bacterium Streptomyces avidinii. Streptavidin homo-tetramers have an extraordinarily high affinity for biotin. With a dissociation constant (Kd) on the order of ≈10−14 mol/L, the binding of biotin to streptavidin is one of the strongest non-covalent interactions known in nature. Streptavidin is used extensively in molecular biology and bionanotechnology due to the streptavidin-biotin complex's resistance to organic solvents, denaturants, detergents, proteolytic enzymes, and extremes of temperature and pH.
A tetrameric protein is a protein with a quaternary structure of four subunits (tetrameric). Homotetramers have four identical subunits, and heterotetramers are complexes of different subunits. A tetramer can be assembled as dimer of dimers with two homodimer subunits, or two heterodimer subunits.

An isopeptide bond is a type of amide bond formed between a carboxyl group of one amino acid and an amino group of another. An isopeptide bond is the linkage between the side chain amino or carboxyl group of one amino acid to the α-carboxyl, α-amino group, or the side chain of another amino acid. In a typical peptide bond, also known as eupeptide bond, the amide bond always forms between the α-carboxyl group of one amino acid and the α-amino group of the second amino acid. Isopeptide bonds are rarer than regular peptide bonds. Isopeptide bonds lead to branching in the primary sequence of a protein. Proteins formed from normal peptide bonds typically have a linear primary sequence.

mRNA display is a display technique used for in vitro protein, and/or peptide evolution to create molecules that can bind to a desired target. The process results in translated peptides or proteins that are associated with their mRNA progenitor via a puromycin linkage. The complex then binds to an immobilized target in a selection step. The mRNA-protein fusions that bind well are then reverse transcribed to cDNA and their sequence amplified via a polymerase chain reaction. The result is a nucleotide sequence that encodes a peptide with high affinity for the molecule of interest.

Fusion proteins or chimeric (kī-ˈmir-ik) proteins are proteins created through the joining of two or more genes that originally coded for separate proteins. Translation of this fusion gene results in a single or multiple polypeptides with functional properties derived from each of the original proteins. Recombinant fusion proteins are created artificially by recombinant DNA technology for use in biological research or therapeutics. Chimeric or chimera usually designate hybrid proteins made of polypeptides having different functions or physico-chemical patterns. Chimeric mutant proteins occur naturally when a complex mutation, such as a chromosomal translocation, tandem duplication, or retrotransposition creates a novel coding sequence containing parts of the coding sequences from two different genes. Naturally occurring fusion proteins are commonly found in cancer cells, where they may function as oncoproteins. The bcr-abl fusion protein is a well-known example of an oncogenic fusion protein, and is considered to be the primary oncogenic driver of chronic myelogenous leukemia.
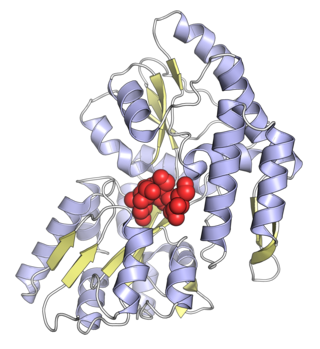
Maltose-binding protein (MBP) is a part of the maltose/maltodextrin system of Escherichia coli, which is responsible for the uptake and efficient catabolism of maltodextrins. It is a complex regulatory and transport system involving many proteins and protein complexes. MBP has an approximate molecular mass of 42.5 kilodaltons.

Meir Wilchek is an Israeli biochemist. He is a professor at the Weizmann Institute of Science.
The Strep-tag system is a method which allows the purification and detection of proteins by affinity chromatography. The Strep-tag II is a synthetic peptide consisting of eight amino acids (Trp-Ser-His-Pro-Gln-Phe-Glu-Lys). This peptide sequence exhibits intrinsic affinity towards Strep-Tactin, a specifically engineered streptavidin, and can be N- or C- terminally fused to recombinant proteins. By exploiting the highly specific interaction, Strep-tagged proteins can be isolated in one step from crude cell lysates. Because the Strep-tag elutes under gentle, physiological conditions, it is especially suited for generation of functional proteins.
Affibody molecules are small, robust proteins engineered to bind to a large number of target proteins or peptides with high affinity, imitating monoclonal antibodies, and are therefore a member of the family of antibody mimetics. Affibody molecules are used in biochemical research and are being developed as potential new biopharmaceutical drugs. These molecules can be used for molecular recognition in diagnostic and therapeutic applications.
The Streptavidin-Binding Peptide (SBP)-Tag is a 38-amino acid sequence that may be engineered into recombinant proteins. Recombinant proteins containing the SBP-Tag bind to streptavidin and this property may be utilized in specific purification, detection or immobilization strategies.

FlAsH-EDT2 is an organoarsenic compound with molecular formula C24H18As2O5S4. Its structure is based around a fluorescein core with two 1,3,2-dithiarsolane substituents. It is used in bioanalytical research as a fluorescent label for visualising proteins in living cells. FlAsH-EDT2 is an abbreviation for fluorescin arsenical hairpin binder-ethanedithiol, and is a pale yellow or pinkish fluorogenic solid. It has a semi-structural formula (C2H4AsS2)2-(C13H5O3)-C6H4COOH, representing the dithiarsolane substituents bound to the hydroxyxanthone core, attached to an o-substituted molecule of benzoic acid.

IBA Lifesciences is a biotechnology company providing products and custom specific services for life science applications in academia and industry worldwide. IBA focusses on two business segments: cell selection and protein purification.
A Spot-tag is a 12-amino acid peptide tag recognized by a single-domain antibody. Due to the small size of a Spot-tag and the robust Spot-nanobody that specifically binds to Spot-tagged proteins, Spot-tag can be used for multiple capture and detection applications: Immunoprecipitation, affinity purification, immunofluorescence, and super-resolution microscopy. Recombinant proteins can be engineered to express the Spot-tag.
The SpyTag/SpyCatcher system is a technology for irreversible conjugation of recombinant proteins. The peptide SpyTag spontaneously reacts with the protein SpyCatcher to form an intermolecular isopeptide bond between the pair. DNA sequence encoding either SpyTag or SpyCatcher can be recombinantly introduced into the DNA sequence encoding a protein of interest, forming a fusion protein. These fusion proteins can be covalently linked when mixed in a reaction through the SpyTag/SpyCatcher system.
References
- ↑ Mahmoudi Gomari, Mohammad; Saraygord-Afshari, Neda; Farsimadan, Marziye; Rostami, Neda; Aghamiri, Shahin; Farajollahi, Mohammad M. (December 2020). "Opportunities and challenges of the tag-assisted protein purification techniques: Applications in the pharmaceutical industry". Biotechnology Advances. 45: 107653. doi:10.1016/j.biotechadv.2020.107653. ISSN 0734-9750. PMID 33157154. S2CID 226276355.
- 1 2 Schmidt, Thomas G.M.; Koepke, Jürgen; Frank, Ronald; Skerra, Arne (1996). "Molecular Interaction Between the Strep-tag Affinity Peptide and its Cognate Target, Streptavidin". Journal of Molecular Biology. 255 (5): 753–66. doi:10.1006/jmbi.1996.0061. PMID 8636976.
- 1 2 Fairhead M, Krndija D, Lowe ED, Howarth M (January 2014). "Plug-and-play pairing via defined divalent streptavidins". Journal of Molecular Biology. 426 (1): 199–214. doi:10.1016/j.jmb.2013.09.016. PMC 4047826 . PMID 24056174.
- 1 2 Zhang, Jin; Campbell, Robert; Ting, Alice; Tsien, Roger (2002). "Creating new fluorescent probes for cell biology". Nat Rev Mol Cell Biol. 3 (12): 906–918. doi:10.1038/nrm976. PMID 12461557. S2CID 11588100.
- ↑ Götzke, Hansjörg; Kilisch, Markus; Martínez-Carranza, Markel; Sograte-Idrissi, Shama; Rajavel, Abirami; Schlichthaerle, Thomas; Engels, Niklas; Jungmann, Ralf; Stenmark, Pål; Opazo, Felipe; Frey, Steffen (2019). "The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications". Nature Communications. 10 (1): 4403. Bibcode:2019NatCo..10.4403G. doi:10.1038/s41467-019-12301-7. PMC 6764986 . PMID 31562305.
- ↑ De Genst, Erwin J.; Guilliams, Tim; Wellens, Joke; O'Day, Elizabeth M.; Waudby, Christopher A.; Meehan, Sarah; Dumoulin, Mireille; Hsu, Shang-Te Danny; Cremades, Nunilo; Verschueren, Koen H.G.; Pardon, Els; Wyns, Lode; Steyaert, Jan; Christodoulou, John; Dobson, Christopher M. (September 2010). "Structure and Properties of a Complex of α-Synuclein and a Single-Domain Camelid Antibody" (PDF). Journal of Molecular Biology. 402 (2): 326–343. doi:10.1016/j.jmb.2010.07.001. PMID 20620148.
- ↑ "CaptureSelect C-tag Affinity Matrix - Thermo Fisher Scientific". www.thermofisher.com.
- ↑ Cooper, Merideth A.; Taris, Joseph E.; Shi, Changhua; Wood, David W. (2018). "A Convenient Split‐Intein Tag Method for the Purification of Tagless Target Proteins". Current Protocols in Protein Science. 91 (1): 5.29.1–5.29.23. doi:10.1002/cpps.46. ISSN 1934-3655. PMID 29516483. S2CID 3749506.
- ↑ Prabhala, Sai Vivek; Gierach, Izabela; Wood, David W. (2022). "The Evolution of Intein-Based Affinity Methods as Reflected in 30 years of Patent History". Frontiers in Molecular Biosciences. 9: 857566. doi: 10.3389/fmolb.2022.857566 . ISSN 2296-889X. PMC 9033041 . PMID 35463948.
- ↑ "iCapTag™ Technology - Protein Capture Science". www.proteincapturescience.com.
- ↑ Einhauer, A.; Jungbauer, A. (2001). "The FLAG™ peptide, a versatile fusion tag for the purification of recombinant proteins". Journal of Biochemical and Biophysical Methods. 49 (1–3): 455–65. doi:10.1016/S0165-022X(01)00213-5. PMID 11694294.
- ↑ Prakriya, Murali; Feske, Stefan; Gwack, Yousang; Srikanth, Sonal; Rao, Anjana; Hogan, Patrick G. (2006). "Orai1 is an essential pore subunit of the CRAC channel". Nature. 443 (7108): 230–3. Bibcode:2006Natur.443..230P. doi:10.1038/nature05122. PMID 16921383. S2CID 4310221.
- ↑ Brune, Karl D.; Liekniņa, Ilva; Sutov, Grigorij; Morris, Alexander R.; Jovicevic, Dejana; Kalniņš, Gints; Kazāks, Andris; Kluga, Rihards; Kastaljana, Sabine; Zajakina, Anna; Jansons, Juris; Skrastiņa, Dace; Spunde, Karīna; Cohen, Alexander A.; Bjorkman, Pamela J.; Morris, Howard R.; Suna, Edgars; Tārs, Kaspars (2021-11-16). "N-Terminal Modification of Gly-His-Tagged Proteins with Azidogluconolactone". ChemBioChem. 22 (22): 3199–3207. doi:10.1002/cbic.202100381. ISSN 1439-7633. PMID 34520613. S2CID 237515136.
- ↑ Ho, Philip WL.; Tse, Zero HM.; Liu, HF.; Lu, S.; Ho, Jessica WM.; Kung, Michelle HW.; Ramsden, David B.; Ho, SL. (2013). "Assessment of cellular estrogenic activity based on estrogen receptor-mediated reduction of soluble-form catechol-O-methyltransferase (COMT) expression in an ELISA-based system". PLOS ONE. 8 (9): e74065. Bibcode:2013PLoSO...874065H. doi: 10.1371/journal.pone.0074065 . PMC 3765251 . PMID 24040167.
- ↑ Keefe, Anthony D.; Wilson, David S.; Seelig, Burckhard; Szostak, Jack W. (2001). "One-Step Purification of Recombinant Proteins Using a Nanomolar-Affinity Streptavidin-Binding Peptide, the SBP-Tag". Protein Expression and Purification. 23 (3): 440–6. doi:10.1006/prep.2001.1515. PMID 11722181.
- ↑ Gelerter, Bruce (June 11, 2014). "PEMF For Treatment Of Corneal Disorders And Injuries".
- ↑ "Epitope Tags & Fusion Proteins – antibodies-online". www.antibodies-online.com.
- ↑ McNutt, Markey C.; Lagace, Thomas A.; Horton, Jay D. (2007). "Catalytic Activity is Not Required for Secreted PCSK9 to Reduce Low Density Lipoprotein Receptors in HepG2 Cells". Journal of Biological Chemistry. 282 (29): 20799–803. doi: 10.1074/jbc.C700095200 . PMID 17537735.
- ↑ Zakeri, Bijan; Howarth, Mark (2010). "Spontaneous Intermolecular Amide Bond Formation between Side Chains for Irreversible Peptide Targeting". Journal of the American Chemical Society. 132 (13): 4526–7. doi:10.1021/ja910795a. PMID 20235501.
- ↑ Zakeri, Bijan; Fierer, Jacob O.; Celik, Emrah; Chittock, Emily C.; Schwarz-Linek, Ulrich; Moy, Vincent T.; Howarth, Mark (2012). "Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin". Proceedings of the National Academy of Sciences. 109 (12): E690–7. Bibcode:2012PNAS..109E.690Z. doi: 10.1073/pnas.1115485109 . PMC 3311370 . PMID 22366317.
- ↑ Veggiani, Gianluca; Nakamura, Tomohiko; Brenner, Michael; Gayet, Raphael; Yan, Jun; Robinson, Carol; Howarth, Mark (2016). "Programmable polyproteams built using twin peptide superglues". Proceedings of the National Academy of Sciences. 113 (5): 1202–7. Bibcode:2016PNAS..113.1202V. doi: 10.1073/pnas.1519214113 . PMC 4747704 . PMID 26787909.
- 1 2 Buldun, Can M.; Jean, Jisoo X.; Bedford, Michael R.; Howarth, Mark (14 February 2018). "SnoopLigase Catalyzes Peptide–Peptide Locking and Enables Solid-Phase Conjugate Isolation". Journal of the American Chemical Society. 140 (8): 3008–3018. doi:10.1021/jacs.7b13237. PMID 29402082. S2CID 207189163.
- ↑ Keeble, Anthony H.; Yadav, Vikash K.; Ferla, Matteo P.; Bauer, Claudia C.; Chuntharpursat-Bon, Eulashini; Huang, Jin; Bon, Robin S.; Howarth, Mark (July 2021). "DogCatcher allows loop-friendly protein-protein ligation". Cell Chemical Biology. 29 (2): 339–350.e10. doi: 10.1016/j.chembiol.2021.07.005 . ISSN 2451-9456. PMC 8878318 . PMID 34324879.
- ↑ Tan, Lee Ling; Hoon, Shawn S.; Wong, Fong T.; Ahmed, S. Ashraf (26 October 2016). "Kinetic Controlled Tag-Catcher Interactions for Directed Covalent Protein Assembly". PLOS ONE. 11 (10): e0165074. Bibcode:2016PLoSO..1165074T. doi: 10.1371/journal.pone.0165074 . PMC 5082641 . PMID 27783674.
- ↑ Ciulli, Bond; Alessi, Craigon (Oct 2021). "Development of BromoTag: A "Bump-and-Hole"–PROTAC System to Induce Potent, Rapid, and Selective Degradation of Tagged Target Proteins". J Med Chem. 64 (20): 15477–15502. doi: 10.1021/acs.jmedchem.1c01532 . PMC 8558867 . PMID 34652918.
- ↑ Chow LT, Vassylyev DG. Application of a Novel CL7/Im7 Affinity System in Purification of Complex and Pharmaceutical Proteins. Methods Mol Biol. 2022;2466:61-82. doi: 10.1007/978-1-0716-2176-9_5. PMID: 35585311.
- ↑ Bedouelle, Hugues; Duplay, Pascale (Feb 1988). "Production in Escherichia coli and one-step purification of bifunctional hybrid proteins which bind maltose. Export of the Klenow polymerase into the periplasmic space". Eur J Biochem. 171 (3): 541–549. doi: 10.1111/j.1432-1033.1988.tb13823.x . PMID 3278900.
- ↑ Minde, David P; Halff, Els F; Tans, Sander (2013). "Designing disorder: Tales of the unexpected tails". Intrinsically Disordered Proteins. 1 (1): 5–15. doi:10.4161/idp.26790. PMC 5424805 . PMID 28516025.
- ↑ Kriznik, Alexandre; Yéléhé-Okouma, Mélissa; Lec, Jean-Christophe; Groshenry, Guillaume; Le Cordier, Hélène; Charron, Christophe; Quinternet, Marc; Mazon, Hortense; Talfournier, François; Boschi-Muller, Sandrine; Jouzeau, Jean-Yves; Reboul, Pascal (Oct 2018). "CRDSAT Generated by pCARGHO: A New Efficient Lectin-Based Affinity Tag Method for Safe, Simple, and Low-Cost Protein Purification". Biotechnol J. 14 (4): 1800214. doi:10.1002/biot.201800214. PMID 30298550. S2CID 52942568.
- 1 2 Schwinn, Marie K.; Machleidt, Thomas; Zimmerman, Kris; Eggers, Christopher T.; Dixon, Andrew S.; Hurst, Robin; Hall, Mary P.; Encell, Lance P.; Binkowski, Brock F.; Wood, Keith V. (2018-02-16). "CRISPR-Mediated Tagging of Endogenous Proteins with a Luminescent Peptide". ACS Chemical Biology. 13 (2): 467–474. doi: 10.1021/acschembio.7b00549 . ISSN 1554-8929. PMID 28892606.