P-bodies, or processing bodies are distinct foci formed by phase separation within the cytoplasm of the eukaryotic cell consisting of many enzymes involved in mRNA turnover. P-bodies are highly conserved structures and have been observed in somatic cells originating from vertebrates and invertebrates, plants and yeast. To date, P-bodies have been demonstrated to play fundamental roles in general mRNA decay, nonsense-mediated mRNA decay, adenylate-uridylate-rich element mediated mRNA decay, and microRNA (miRNA) induced mRNA silencing. Not all mRNAs which enter P-bodies are degraded, as it has been demonstrated that some mRNAs can exit P-bodies and re-initiate translation. Purification and sequencing of the mRNA from purified processing bodies showed that these mRNAs are largely translationally repressed upstream of translation initiation and are protected from 5' mRNA decay.

The 5' cap of eukaryotic messenger RNA is bound at all times by various cap-binding complexes (CBCs).

5′-3′ exoribonuclease 1 (Xrn1) is a protein that in humans is encoded by the XRN1 gene. Xrn1 hydrolyses RNA in the 5′ to 3′ direction.

RNA-binding protein 8A is a protein that in humans is encoded by the RBM8A gene.
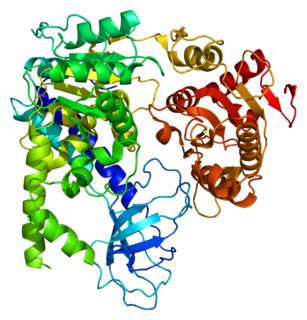
Regulator of nonsense transcripts 1 is a protein that in humans is encoded by the UPF1 gene.
Probable ATP-dependent RNA helicase DDX5 also known as DEAD box protein 5 or RNA helicase p68 is an enzyme that in humans is encoded by the DDX5 gene.

Regulator of nonsense transcripts 2 is a protein that in humans is encoded by the UPF2 gene.

Trinucleotide repeat-containing gene 6A protein is a protein that in humans is encoded by the TNRC6A gene.

Regulator of nonsense transcripts 3B is a protein that in humans is encoded by the UPF3B gene.

mRNA-decapping enzyme 2 is a protein that in humans is encoded by the DCP2 gene.

Serine/threonine-protein kinase SMG1 is an enzyme that in humans is encoded by the SMG1 gene. SMG1 belongs to the phosphatidylinositol 3-kinase-related kinase protein family.

Far upstream element-binding protein 2 is a protein that in humans is encoded by the KHSRP gene.

Regulator of nonsense transcripts 3A is a protein that in humans is encoded by the UPF3A gene.

Scavenger mRNA-decapping enzyme DcpS is a protein that in humans is encoded by the DCPS gene.

Enhancer of mRNA-decapping protein 3 is a protein that in humans is encoded by the EDC3 gene.

mRNA-decapping enzyme 1B is a protein that in humans is encoded by the DCP1B gene.
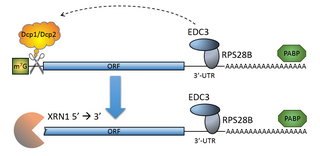
The process of messenger RNA decapping consists of hydrolysis of the 5' cap structure on the RNA exposing a 5' monophosphate. In eukaryotes, this 5' monophosphate is a substrate for the 5' exonuclease Xrn1 and the mRNA is quickly destroyed. There are many situations which may lead to the removal of the cap, some of which are discussed below.

The mRNA decapping complex is a protein complex in eukaryotic cells responsible for removal of the 5' cap. The active enzyme of the decapping complex is the bilobed Nudix family enzyme Dcp2, which hydrolyzes 5' cap and releases 7mGDP and a 5'-monophosphorylated mRNA. This decapped mRNA is inhibited for translation and will be degraded by exonucleases. The core decapping complex is conserved in eukaryotes. Dcp2 is activated by Decapping Protein 1 (Dcp1) and in higher eukaryotes joined by the scaffold protein VCS. Together with many other accessory proteins, the decapping complex assembles in P-bodies in the cytoplasm.
Cryptic unstable transcripts (CUTs) are a subset of non-coding RNAs (ncRNAs) that are produced from intergenic and intragenic regions. CUTs were first observed in S. cerevisiae yeast models and are found in most eukaryotes. Some basic characteristics of CUTs include a length of around 200–800 base pairs, a 5' cap, poly-adenylated tail, and rapid degradation due to the combined activity of poly-adenylating polymerases and exosome complexes. CUT transcription occurs through RNA Polymerase II and initiates from nucleosome-depleted regions, often in an antisense orientation. To date, CUTs have a relatively uncharacterized function but have been implicated in a number of putative gene regulation and silencing pathways. Thousands of loci leading to the generation of CUTs have been described in the yeast genome. Additionally, stable uncharacterized transcripts, or SUTs, have also been detected in cells and bear many similarities to CUTs but are not degraded through the same pathways.
M7GpppN-mRNA hydrolase (EC 3.6.1.62, DCP2, NUDT16, D10 protein, D9 protein, D10 decapping enzyme, decapping enzyme) is an enzyme with systematic name m7GpppN-mRNA m7GDP phosphohydrolase. This enzyme catalyses the following chemical reaction