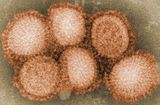
Influenza A virus (IAV) is a pathogen that causes the flu in birds and some mammals, including humans. It is an RNA virus whose subtypes have been isolated from wild birds. Occasionally, it is transmitted from wild to domestic birds, and this may cause severe disease, outbreaks, or human influenza pandemics.

Avian influenza, also known as avian flu, is a bird flu caused by the influenza A virus, which can infect people. It is similar to other types of animal flu in that it is caused by a virus strain that has adapted to a specific host. The type with the greatest risk is highly pathogenic avian influenza (HPAI).

Orthomyxoviridae is a family of negative-sense RNA viruses. It includes seven genera: Alphainfluenzavirus, Betainfluenzavirus, Gammainfluenzavirus, Deltainfluenzavirus, Isavirus, Thogotovirus, and Quaranjavirus. The first four genera contain viruses that cause influenza in birds and mammals, including humans. Isaviruses infect salmon; the thogotoviruses are arboviruses, infecting vertebrates and invertebrates. The Quaranjaviruses are also arboviruses, infecting vertebrates (birds) and invertebrates (arthropods).

Influenza A virus subtype H5N1 (A/H5N1) is a subtype of the influenza A virus which can cause illness in humans and many other species. A bird-adapted strain of H5N1, called HPAI A(H5N1) for highly pathogenic avian influenza virus of type A of subtype H5N1, is the highly pathogenic causative agent of H5N1 flu, commonly known as avian influenza. It is enzootic in many bird populations, especially in Southeast Asia. One strain of HPAI A(H5N1) is spreading globally after first appearing in Asia. It is epizootic and panzootic, killing tens of millions of birds and spurring the culling of hundreds of millions of others to stem its spread. Many references to "bird flu" and H5N1 in the popular media refer to this strain.

Swine influenza is an infection caused by any of several types of swine influenza viruses. Swine influenza virus (SIV) or swine-origin influenza virus (S-OIV) refers to any strain of the influenza family of viruses that is endemic in pigs. As of 2009, identified SIV strains include influenza C and the subtypes of influenza A known as H1N1, H1N2, H2N1, H3N1, H3N2, and H2N3.

An influenza pandemic is an epidemic of an influenza virus that spreads across a large region and infects a large proportion of the population. There have been six major influenza epidemics in the last 140 years, with the 1918 flu pandemic being the most severe; this is estimated to have been responsible for the deaths of 50–100 million people. The 2009 swine flu pandemic resulted in under 300,000 deaths and is considered relatively mild. These pandemics occur irregularly.

Influenza A virus subtype H3N2 (A/H3N2) is a subtype of viruses that causes influenza (flu). H3N2 viruses can infect birds and mammals. In birds, humans, and pigs, the virus has mutated into many strains. In years in which H3N2 is the predominant strain, there are more hospitalizations.

The global spread of H5N1 influenza in birds is considered a significant pandemic threat. While other H5N1 influenza strains are known, they are significantly different on a genetic level from a recent, highly pathogenic, emergent strain of H5N1, which was able to achieve hitherto unprecedented global spread in 2008. The H5N1 strain is a fast-mutating, highly pathogenic avian influenza virus (HPAI) found in multiple bird species. It is both epizootic and panzootic. Unless otherwise indicated, "H5N1" in this timeline refers to the recent highly pathogenic strain of H5N1.

Transmission and infection of H5N1 from infected avian sources to humans has been a concern since the first documented case of human infection in 1997, due to the global spread of H5N1 that constitutes a pandemic threat.

H5N1 genetic structure is the molecular structure of the H5N1 virus's RNA.

H5N1 clinical trials are clinical trials concerning H5N1 vaccines, which are intended to provide immunization to influenza A virus subtype H5N1. They are intended to discover pharmacological effects and identify any adverse reactions the vaccines may achieve in humans.

Spanish flu research concerns studies regarding the causes and characteristics of the Spanish flu, a variety of influenza that in 1918 was responsible for the worst influenza pandemic in modern history. Many theories about the origins and progress of the Spanish flu persisted in the literature, but it was not until 2005, when various samples of lung tissue were recovered from American World War I soldiers and from an Inupiat woman buried in permafrost in a mass grave in Brevig Mission, Alaska, that significant genetic research was made possible.

Steven Lloyd Salzberg is an American computational biologist and computer scientist who is a Bloomberg Distinguished Professor of Biomedical Engineering, Computer Science, and Biostatistics at Johns Hopkins University, where he is also Director of the Center for Computational Biology.

The global spread of H5N1 in birds is considered a significant pandemic threat.

Fujian flu refers to flu caused by either a Fujian human flu strain of the H3N2 subtype of the Influenza A virus or a Fujian bird flu strain of the H5N1 subtype of the Influenza A virus. These strains are named after Fujian, a coastal province in Southeast China.

The Goose Guangdong virus refers to the strain A/Goose/Guangdong/1/96 (Gs/Gd)-like H5N1 HPAI viruses. It is a strain of the Influenzavirus A subtype H5N1 virus that was first detected in a goose in Guangdong in 1996. It is an HPAI virus, meaning that it can kill a very high percentage of chickens in a flock in mere days. It is believed to be the immediate precursor of the current dominant strain of HPAI A(H5N1) that evolved from 1999 to 2002 creating the Z genotype that is spreading globally and is epizootic and panzootic, killing tens of millions of birds and spurring the culling of hundreds of millions of others to stem its spread.
Jeffery K. Taubenberger is an American virologist. With Ann Reid, he was the first to sequence the genome of the influenza virus which caused the 1918 pandemic of Spanish flu. He is Chief of the Viral Pathogenesis and Evolution Section, Laboratory of Infectious Diseases, National Institute of Allergy and Infectious Diseases, National Institutes of Health. Taubenberger's laboratory studies a number of viruses, including influenza A viruses (IAVs), which are the pathogens that cause yearly flu epidemics and have caused periodic pandemics, such as the 1968 outbreak that killed an estimated one million people. His research aims to inform public health strategies on several important aspects of flu: seasonal flu; avian flu, which circulates among birds and has infected humans in the past; swine flu, which circulates among pigs and has infected humans in the past; and pandemic flu, which can arise from numerous sources and spread quickly because humans have little to no immunity to it.

The pandemic H1N1/09 virus is a swine origin influenza A virus subtype H1N1 strain that was responsible for the 2009 swine flu pandemic. This strain is often called swine flu by the public media. For other names, see the Nomenclature section below.

Elodie Ghedin is a Canadian parasitologist and virologist as well as a professor at the New York University Center for Genomics and Systems Biology. Her work focuses on the molecular biology and genomics of the parasites that cause diseases such as elephantiasis, and river blindness, and on the evolution of the influenza virus. She was named a 2011 MacArthur Fellow, a 2012 Kavli Frontier of Science Fellow, and a 2017 American Academy of Microbiology Fellow. She also was Awarded the Chancellor’s Distinguished Research Award in 2010.