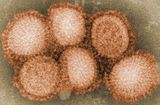
Influenza A virus (IAV) is a pathogen that causes the flu in birds and some mammals, including humans. It is an RNA virus whose subtypes have been isolated from wild birds. Occasionally, it is transmitted from wild to domestic birds, and this may cause severe disease, outbreaks, or human influenza pandemics.

Antigenic shift is the process by which two or more different strains of a virus, or strains of two or more different viruses, combine to form a new subtype having a mixture of the surface antigens of the two or more original strains. The term is often applied specifically to influenza, as that is the best-known example, but the process is also known to occur with other viruses, such as visna virus in sheep. Antigenic shift is a specific case of reassortment or viral shift that confers a phenotypic change.

Influenza A virus subtype H5N1 (A/H5N1) is a subtype of the influenza A virus which can cause illness in humans and many other species. A bird-adapted strain of H5N1, called HPAI A(H5N1) for highly pathogenic avian influenza virus of type A of subtype H5N1, is the highly pathogenic causative agent of H5N1 flu, commonly known as avian influenza. It is enzootic in many bird populations, especially in Southeast Asia. One strain of HPAI A(H5N1) is spreading globally after first appearing in Asia. It is epizootic and panzootic, killing tens of millions of birds and spurring the culling of hundreds of millions of others to stem its spread. Many references to "bird flu" and H5N1 in the popular media refer to this strain.

Influenza vaccines, also known as flu shots, are vaccines that protect against infection by influenza viruses. New versions of the vaccines are developed twice a year, as the influenza virus rapidly changes. While their effectiveness varies from year to year, most provide modest to high protection against influenza. The United States Centers for Disease Control and Prevention (CDC) estimates that vaccination against influenza reduces sickness, medical visits, hospitalizations, and deaths. Immunized workers who do catch the flu return to work half a day sooner on average. Vaccine effectiveness in those over 65 years old remains uncertain due to a lack of high-quality research.

The Hong Kong flu, also known as the 1968 flu pandemic, was a flu pandemic whose outbreak in 1968 and 1969 killed between one and four million people globally. It is among the deadliest pandemics in history, and was caused by an H3N2 strain of the influenza A virus. The virus was descended from H2N2 through antigenic shift, a genetic process in which genes from multiple subtypes are reassorted to form a new virus.

Reassortment is the mixing of the genetic material of a species into new combinations in different individuals. Several different processes contribute to reassortment, including assortment of chromosomes, and chromosomal crossover. It is particularly used when two similar viruses that are infecting the same cell exchange genetic material. In particular, reassortment occurs among influenza viruses, whose genomes consist of eight distinct segments of RNA. These segments act like mini-chromosomes, and each time a flu virus is assembled, it requires one copy of each segment.

Swine influenza is an infection caused by any of several types of swine influenza viruses. Swine influenza virus (SIV) or swine-origin influenza virus (S-OIV) refers to any strain of the influenza family of viruses that is endemic in pigs. As of 2009, identified SIV strains include influenza C and the subtypes of influenza A known as H1N1, H1N2, H2N1, H3N1, H3N2, and H2N3.

Flu season is an annually recurring time period characterized by the prevalence of an outbreak of influenza (flu). The season occurs during the cold half of the year in each hemisphere. It takes approximately two days to show symptoms. Influenza activity can sometimes be predicted and even tracked geographically. While the beginning of major flu activity in each season varies by location, in any specific location these minor epidemics usually take about three weeks to reach its pinnacle, and another three weeks to significantly diminish.

In virology, influenza A virus subtype H1N1 (A/H1N1) is a subtype of influenza A virus. Major outbreaks of H1N1 strains in humans include the 1918 Spanish flu pandemic, the 1977 Russian flu pandemic and the 2009 swine flu pandemic. It is an orthomyxovirus that contains the glycoproteins hemagglutinin and neuraminidase. For this reason, they are described as H1N1, H1N2 etc., depending on the type of H or N antigens they express with metabolic synergy. Hemagglutinin causes red blood cells to clump together and binds the virus to the infected cell. Neuraminidase is a type of glycoside hydrolase enzyme which helps to move the virus particles through the infected cell and assist in budding from the host cells.

An influenza pandemic is an epidemic of an influenza virus that spreads across a large region and infects a large proportion of the population. There have been six major influenza epidemics in the last 140 years, with the 1918 flu pandemic being the most severe; this is estimated to have been responsible for the deaths of 50–100 million people. The most recent, the 2009 swine flu pandemic, resulted in under 300,000 deaths and is considered relatively mild. These pandemics occur irregularly.

Influenza B virus is the only species in the genus Betainfluenzavirus in the virus family Orthomyxoviridae.

Fujian flu refers to flu caused by either a Fujian human flu strain of the H3N2 subtype of the Influenza A virus or a Fujian bird flu strain of the H5N1 subtype of the Influenza A virus. These strains are named after Fujian, a coastal province in Southeast China.

Influenza, commonly known as "the flu", is an infectious disease caused by influenza viruses. Symptoms range from mild to severe and often include fever, runny nose, sore throat, muscle pain, headache, coughing, and fatigue. These symptoms begin from one to four days after exposure to the virus and last for about 2–8 days. Diarrhea and vomiting can occur, particularly in children. Influenza may progress to pneumonia, which can be caused by the virus or by a subsequent bacterial infection. Other complications of infection include acute respiratory distress syndrome, meningitis, encephalitis, and worsening of pre-existing health problems such as asthma and cardiovascular disease.

The 2009 swine flu pandemic, caused by the H1N1/swine flu/ influenza virus and declared by the World Health Organization (WHO) from June 2009 to August 2010, is the third recent flu pandemic involving the H1N1 virus. The first two cases were discovered independently in the United States in April 2009. The virus appeared to be a new strain of H1N1 that resulted from a previous triple reassortment of bird, swine, and human flu viruses which further combined with a Eurasian pig flu virus, leading to the term "swine flu".
The 2009 flu pandemic in the United States was caused by a novel strain of the Influenza A/H1N1 virus, commonly referred to as "swine flu," that was first detected on 15 April 2009. While the 2009 H1N1 virus strain was commonly referred to as "swine flu," there is no evidence that it is endemic to pigs or of transmission from pigs to people; instead, the virus spreads from person to person. On April 25, the World Health Organization declared a public health emergency, followed concurringly by the Obama administration on April 26.

This article covers the chronology of the 2009 novel influenza A (H1N1) pandemic. Flag icons denote the first announcements of confirmed cases by the respective nation-states, their first deaths, and relevant sessions and announcements of the World Health Organization (WHO), the European Union , and the U.S. Centers for Disease Control (CDC).

The pandemic H1N1/09 virus is a swine origin influenza A virus subtype H1N1 strain that was responsible for the 2009 swine flu pandemic. This strain is often called swine flu by the public media. For other names, see the Nomenclature section below.

The 2009 swine flu pandemic vaccines were influenza vaccines developed to protect against the pandemic H1N1/09 virus. These vaccines either contained inactivated (killed) influenza virus, or weakened live virus that could not cause influenza. The killed virus was injected, while the live virus was given as a nasal spray. Both these types of vaccine were produced by growing the virus in chicken eggs. Around three billion doses were produced, with delivery in November 2009.

The 2009 flu pandemic was a global outbreak of a new strain of influenza A virus subtype H1N1, first identified in April 2009, termed Pandemic H1N1/09 virus by the World Health Organization (WHO) and colloquially called swine flu. The outbreak was first observed in Mexico, and quickly spread globally. On 11 June 2009, WHO declared the outbreak to be a pandemic. The overwhelming majority of patients experience mild symptoms", but some persons are in higher risk groups, such as those with asthma, diabetes, obesity, heart disease, or who are pregnant or have a weakened immune system. In the rare severe cases, around 3–5 days after symptoms manifest, the person's condition declines quickly, often to the point respiratory failure.
The Centers for Disease Control and Prevention (CDC) estimates that, as of April 4, 2020, the 2019–2020 United States flu season had caused 39 million to 56 million flu illnesses, 410,000 to 740,000 hospitalizations and 24,000 to 62,000 deaths. In January 2020, the Director of the National Institute of Allergies and Infectious Diseases, Dr. Anthony Fauci expected the 2019–2020 flu season to be one of the worst in several years, at least as severe as the 2017–2018 season. By the third week in February the seasonal flu was near its peak with over 26 million people sickened, 250,000 hospitalized, and 14,000 who died. Experts said that the flu came in two waves, with a hard impact on children. The season began in October, earlier than usual, with the expected wave of influenza B virus. The number of children who died, 105, was higher in late February than any season for the past ten years with about 67% associated with influenza B viruses. The second wave came with the influx of influenza A viruses, such as H1N1. According to preliminary burden estimates for the 2019–2020 flu season there were between 39 and 56 million flu cases; 18–26 million doctor visits; 410,000 to 740,000 hospitalizations, and between 24,000 and 62,000 deaths. The unusually abrupt decline in cases by April 2020 was attributed to the effects of widespread social distancing and lockdowns aimed at COVID-19, shortening the influenza season by 5–6 weeks.