Lectins are carbohydrate-binding proteins that are highly specific for sugar groups that are part of other molecules, so cause agglutination of particular cells or precipitation of glycoconjugates and polysaccharides. Lectins have a role in recognition at the cellular and molecular level and play numerous roles in biological recognition phenomena involving cells, carbohydrates, and proteins. Lectins also mediate attachment and binding of bacteria, viruses, and fungi to their intended targets.

Influenza A virus (IAV) is a pathogen that causes the flu in birds and some mammals, including humans. It is an RNA virus whose subtypes have been isolated from wild birds. Occasionally, it is transmitted from wild to domestic birds, and this may cause severe disease, outbreaks, or human influenza pandemics.

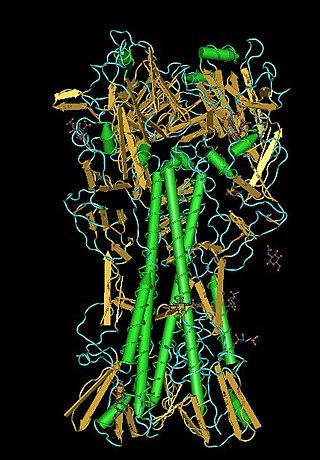
Influenza hemagglutinin (HA) or haemagglutinin[p] is a homotrimeric glycoprotein found on the surface of influenza viruses and is integral to its infectivity.

Orthomyxoviridae is a family of negative-sense RNA viruses. It includes seven genera: Alphainfluenzavirus, Betainfluenzavirus, Gammainfluenzavirus, Deltainfluenzavirus, Isavirus, Thogotovirus, and Quaranjavirus. The first four genera contain viruses that cause influenza in birds and mammals, including humans. Isaviruses infect salmon; the thogotoviruses are arboviruses, infecting vertebrates and invertebrates. The Quaranjaviruses are also arboviruses, infecting vertebrates (birds) and invertebrates (arthropods).
Antigenic drift is a kind of genetic variation in viruses, arising from the accumulation of mutations in the virus genes that code for virus-surface proteins that host antibodies recognize. This results in a new strain of virus particles that is not effectively inhibited by the antibodies that prevented infection by previous strains. This makes it easier for the changed virus to spread throughout a partially immune population. Antigenic drift occurs in both influenza A and influenza B viruses.
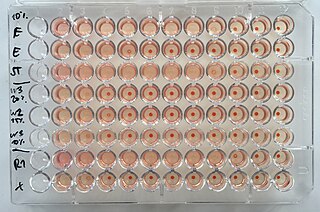

The hemagglutination assay or haemagglutination assay (HA) and the hemagglutination inhibition assay were developed in 1941–42 by American virologist George Hirst as methods for quantifying the relative concentration of viruses, bacteria, or antibodies.
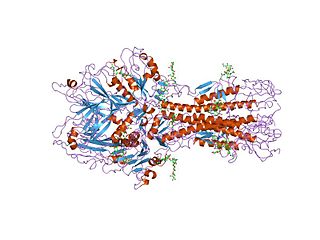
Hemagglutinin esterase (HEs) is a glycoprotein that certain enveloped viruses possess and use as an invading mechanism. HEs helps in the attachment and destruction of certain sialic acid receptors that are found on the host cell surface. Viruses that possess HEs include influenza C virus, toroviruses, and coronaviruses of the subgenus Embecovirus. HEs is a dimer transmembrane protein consisting of two monomers, each monomer is made of three domains. The three domains are: membrane fusion, esterase, and receptor binding domains.
Hemagglutination, or haemagglutination, is a specific form of agglutination that involves red blood cells (RBCs). It has two common uses in the laboratory: blood typing and the quantification of virus dilutions in a haemagglutination assay.

Influenza C virus is the only species in the genus Gammainfluenzavirus, in the virus family Orthomyxoviridae, which like other influenza viruses, causes influenza.

H5N1 genetic structure is the molecular structure of the H5N1 virus's RNA.

A virosome is a drug or vaccine delivery mechanism consisting of unilamellar phospholipid membrane vesicle incorporating virus derived proteins to allow the virosomes to fuse with target cells. Viruses are infectious agents that can replicate in their host organism, however virosomes do not replicate. The properties that virosomes share with viruses are based on their structure; virosomes are essentially safely modified viral envelopes that contain the phospholipid membrane and surface glycoproteins. As a drug or vaccine delivery mechanism they are biologically compatible with many host organisms and are also biodegradable. The use of reconstituted virally derived proteins in the formation of the virosome allows for the utilization of what would otherwise be the immunogenic properties of a live-attenuated virus, but is instead a safely killed virus. A safely killed virus can serve as a promising vector because it won't cause infection and the viral structure allows the virosome to recognize specific components of its target cells.
Antigenic variation or antigenic alteration refers to the mechanism by which an infectious agent such as a protozoan, bacterium or virus alters the proteins or carbohydrates on its surface and thus avoids a host immune response, making it one of the mechanisms of antigenic escape. It is related to phase variation. Antigenic variation not only enables the pathogen to avoid the immune response in its current host, but also allows re-infection of previously infected hosts. Immunity to re-infection is based on recognition of the antigens carried by the pathogen, which are "remembered" by the acquired immune response. If the pathogen's dominant antigen can be altered, the pathogen can then evade the host's acquired immune system. Antigenic variation can occur by altering a variety of surface molecules including proteins and carbohydrates. Antigenic variation can result from gene conversion, site-specific DNA inversions, hypermutation, or recombination of sequence cassettes. The result is that even a clonal population of pathogens expresses a heterogeneous phenotype. Many of the proteins known to show antigenic or phase variation are related to virulence.

In virology, a spike protein or peplomer protein is a protein that forms a large structure known as a spike or peplomer projecting from the surface of an enveloped virus. The proteins are usually glycoproteins that form dimers or trimers.
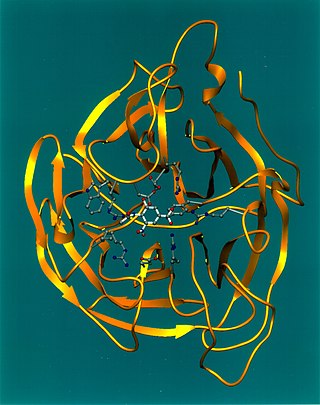
Viral neuraminidase is a type of neuraminidase found on the surface of influenza viruses that enables the virus to be released from the host cell. Neuraminidases are enzymes that cleave sialic acid groups from glycoproteins. Viral neuraminidase was discovered by Alfred Gottschalk at the Walter and Eliza Hall Institute in 1957. Neuraminidase inhibitors are antiviral agents that inhibit influenza viral neuraminidase activity and are of major importance in the control of influenza.
Hemagglutinin-neuraminidase refers to a single viral protein that has both hemagglutinin and (endo) neuraminidase EC 3.2.1.18 activity. This is in contrast to the proteins found in influenza, where both functions exist but in two separate proteins. Its neuraminidase domain has the CAZy designation glycoside hydrolase family 83 (GH83).
FI6 is an antibody that targets a protein found on the surface of all influenza A viruses called hemagglutinin. FI6 is the only known antibody found to bind all 16 subtypes of the influenza A virus hemagglutinin and is hoped to be useful for a universal influenza virus therapy.

George Keble Hirst, M.D. was an American virologist and science administrator who was among the first to study the molecular biology and genetics of animal viruses, especially influenza virus. He directed the Public Health Research Institute in New York City (1956–1981), and was also the founding editor-in-chief of Virology, the first English-language journal to focus on viruses. He is particularly known for inventing the hemagglutination assay, a simple method for quantifying viruses, and adapting it into the hemagglutination inhibition assay, which measures virus-specific antibodies in serum. He was the first to discover that viruses can contain enzymes, and the first to propose that virus genomes can consist of discontinuous segments. The New York Times described him as "a pioneer in molecular virology."
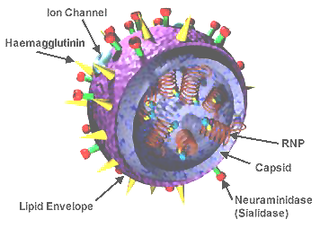
A universal flu vaccine is a flu vaccine that is effective against all influenza strains regardless of the virus sub type, antigenic drift or antigenic shift. Hence it should not require modification from year to year. As of 2021 no universal flu vaccine had been approved for general use, several were in development, and one was in clinical trial.

Influenza D virus is a species in the virus genus Deltainfluenzavirus, in the family Orthomyxoviridae, that causes influenza.
Bat mumps orthorubulavirus, formerly Bat mumps rubulavirus (BMV), is a member of genus Orthorubulavirus, family Paramyxoviridae, and order Mononegavirales. Paramyxoviridae viruses were first isolated from bats using heminested PCR with degenerate primers. This process was then followed by Sanger sequencing. A specific location of this virus is not known because it was isolated from bats worldwide. Although multiple paramyxoviridae viruses have been isolated worldwide, BMV specifically has not been isolated thus far. However, BMV was detected in African fruit bats, but no infectious form has been isolated to date. It is known that BMV is transmitted through saliva in the respiratory system of bats. While the virus was considered its own species for a few years, phylogenetic analysis has since shown that it is a member of Mumps orthorubulavirus.