
A cell wall is a structural layer that surrounds some cell types, found immediately outside the cell membrane. It can be tough, flexible, and sometimes rigid. Primarily, it provides the cell with structural support, shape, protection, and functions as a selective barrier. Another vital role of the cell wall is to help the cell withstand osmotic pressure and mechanical stress. While absent in many eukaryotes, including animals, cell walls are prevalent in other organisms such as fungi, algae and plants, and are commonly found in most prokaryotes, with the exception of mollicute bacteria.
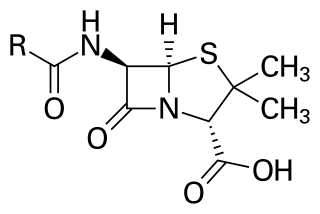
Penicillins are a group of β-lactam antibiotics originally obtained from Penicillium moulds, principally P. chrysogenum and P. rubens. Most penicillins in clinical use are synthesised by P. chrysogenum using deep tank fermentation and then purified. A number of natural penicillins have been discovered, but only two purified compounds are in clinical use: penicillin G and penicillin V. Penicillins were among the first medications to be effective against many bacterial infections caused by staphylococci and streptococci. They are still widely used today for different bacterial infections, though many types of bacteria have developed resistance following extensive use.
Peptidoglycan or murein is a unique large macromolecule, a polysaccharide, consisting of sugars and amino acids that forms a mesh-like peptidoglycan layer (sacculus) that surrounds the bacterial cytoplasmic membrane. The sugar component consists of alternating residues of β-(1,4) linked N-acetylglucosamine (NAG) and N-acetylmuramic acid (NAM). Attached to the N-acetylmuramic acid is an oligopeptide chain made of three to five amino acids. The peptide chain can be cross-linked to the peptide chain of another strand forming the 3D mesh-like layer. Peptidoglycan serves a structural role in the bacterial cell wall, giving structural strength, as well as counteracting the osmotic pressure of the cytoplasm. This repetitive linking results in a dense peptidoglycan layer which is critical for maintaining cell form and withstanding high osmotic pressures, and it is regularly replaced by peptidoglycan production. Peptidoglycan hydrolysis and synthesis are two processes that must occur in order for cells to grow and multiply, a technique carried out in three stages: clipping of current material, insertion of new material, and re-crosslinking of existing material to new material.
Autolysins are endogenous lytic enzymes that break down the peptidoglycan components of biological cells which enables the separation of daughter cells following cell division. They are involved in cell growth, cell wall metabolism, cell division and separation, as well as peptidoglycan turnover and have similar functions to lysozymes.
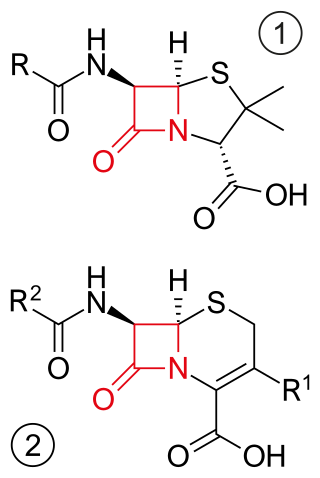
β-lactam antibiotics are antibiotics that contain a beta-lactam ring in their chemical structure. This includes penicillin derivatives (penams), cephalosporins and cephamycins (cephems), monobactams, carbapenems and carbacephems. Most β-lactam antibiotics work by inhibiting cell wall biosynthesis in the bacterial organism and are the most widely used group of antibiotics. Until 2003, when measured by sales, more than half of all commercially available antibiotics in use were β-lactam compounds. The first β-lactam antibiotic discovered, penicillin, was isolated from a strain of Penicillium rubens.

Lysozyme is an antimicrobial enzyme produced by animals that forms part of the innate immune system. It is a glycoside hydrolase that catalyzes the following process:
DD-transpeptidase is a bacterial enzyme that catalyzes the transfer of the R-L-αα-D-alanyl moiety of R-L-αα-D-alanyl-D-alanine carbonyl donors to the γ-OH of their active-site serine and from this to a final acceptor. It is involved in bacterial cell wall biosynthesis, namely, the transpeptidation that crosslinks the peptide side chains of peptidoglycan strands.
Penicillin-binding proteins (PBPs) are a group of proteins that are characterized by their affinity for and binding of penicillin. They are a normal constituent of many bacteria; the name just reflects the way by which the protein was discovered. All β-lactam antibiotics bind to PBPs, which are essential for bacterial cell wall synthesis. PBPs are members of a subgroup of enzymes called transpeptidases. Specifically, PBPs are DD-transpeptidases.
The bacterium, despite its simplicity, contains a well-developed cell structure which is responsible for some of its unique biological structures and pathogenicity. Many structural features are unique to bacteria and are not found among archaea or eukaryotes. Because of the simplicity of bacteria relative to larger organisms and the ease with which they can be manipulated experimentally, the cell structure of bacteria has been well studied, revealing many biochemical principles that have been subsequently applied to other organisms.

Lysins, also known as endolysins or murein hydrolases, are hydrolytic enzymes produced by bacteriophages in order to cleave the host's cell wall during the final stage of the lytic cycle. Lysins are highly evolved enzymes that are able to target one of the five bonds in peptidoglycan (murein), the main component of bacterial cell walls, which allows the release of progeny virions from the lysed cell. Cell-wall-containing Archaea are also lysed by specialized pseudomurein-cleaving lysins, while most archaeal viruses employ alternative mechanisms. Similarly, not all bacteriophages synthesize lysins: some small single-stranded DNA and RNA phages produce membrane proteins that activate the host's autolytic mechanisms such as autolysins.
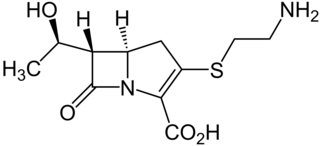
Thienamycin is one of the most potent naturally produced antibiotics known thus far, discovered in Streptomyces cattleya in 1976. Thienamycin has excellent activity against both Gram-positive and Gram-negative bacteria and is resistant to bacterial β-lactamase enzymes. Thienamycin is a zwitterion at pH 7.

Peptidoglycan recognition protein 2(PGLYRP2) is an enzyme, N-acetylmuramoyl-L-alanine amidase (NAMLAA), that hydrolyzes bacterial cell wall peptidoglycan and is encoded by the PGLYRP2 gene.
Cephalosporins are a broad class of bactericidal antibiotics that include the β-lactam ring and share a structural similarity and mechanism of action with other β-lactam antibiotics. The cephalosporins have the ability to kill bacteria by inhibiting essential steps in the bacterial cell wall synthesis which in the end results in osmotic lysis and death of the bacterial cell. Cephalosporins are widely used antibiotics because of their clinical efficiency and desirable safety profile.

Glucanases are enzymes that break down large polysaccharides via hydrolysis. The product of the hydrolysis reaction is called a glucan, a linear polysaccharide made of up to 1200 glucose monomers, held together with glycosidic bonds. Glucans are abundant in the endosperm cell walls of cereals such as barley, rye, sorghum, rice, and wheat. Glucanases are also referred to as lichenases, hydrolases, glycosidases, glycosyl hydrolases, and/or laminarinases. Many types of glucanases share similar amino acid sequences but vastly different substrates. Of the known endo-glucanases, 1,3-1,4-β-glucanase is considered the most active.

Otto Kandler was a German botanist and microbiologist. Until his retirement in 1986 he was professor of botany at the Ludwig Maximilian University of Munich.

Lipid II is a precursor molecule in the synthesis of the cell wall of bacteria. It is a peptidoglycan, which is amphipathic and named for its bactoprenol hydrocarbon chain, which acts as a lipid anchor, embedding itself in the bacterial cell membrane. Lipid II must translocate across the cell membrane to deliver and incorporate its disaccharide-pentapeptide "building block" into the peptidoglycan mesh. Lipid II is the target of several antibiotics.
The bacterial murein precursor exporter (MPE) family is a member of the cation diffusion facilitator (CDF) superfamily of membrane transporters. Members of the MPE family are found in a large variety of Gram-negative and Gram-positive bacteria and facilitate the translocation of lipid-linked murein precursors. A representative list of proteins belonging to the MPE family can be found in the Transporter Classification Database.
OBPgp279 is an endolysin that hydrolyzes peptidoglycan, a major constituent in bacterial membrane. OBPgp279 is found in Pseudomonas fluorescens phage OBP, which belongs in the Myoviridae family of bacteriophages. Because of its role in hydrolyzing the peptidoglycan layer, OBPgp279 is a key enzyme in the lytic cycle of the OBP bacteriophage; it allows the bacteriophage to lyse its host internally to escape. Unlike other endolysins, OBPgp279 does not rely on holins to perforate the inner bacterial membrane in order to reach the peptidoglycan layer. Although OBPgp279 is not a well-studied enzyme, it has garnered interest as a potential antibacterial protein due to its activity against multidrug-resistant gram-negative bacteria.

Peptidoglycan recognition protein 3 is an antibacterial and anti-inflammatory innate immunity protein that in humans is encoded by the PGLYRP3 gene.
Ulvan lyase is an enzyme found within the cell-wall of the marine organisms of the genus Ulva (algae), and some marine bacteria. A lyase is a class of enzyme that catalyzes the breakdown of chemical bonds through an elimination reaction mechanism, rather than a substitution reaction mechanism. Ulvan lyase belongs to the polysaccharide lyase family, a type of enzyme that primarily functions to cleave glycosidic linkages in polysaccharides.














