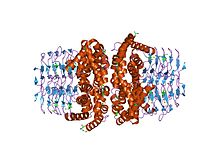
Cysteine is a semiessential proteinogenic amino acid with the formula HOOC−CH(−NH2)−CH2−SH. The thiol side chain in cysteine often participates in enzymatic reactions as a nucleophile. Cysteine is chiral, with only L-cysteine being found in nature.

Methionine is an essential amino acid in humans.
In molecular biology, biosynthesis is a multi-step, enzyme-catalyzed process where substrates are converted into more complex products in living organisms. In biosynthesis, simple compounds are modified, converted into other compounds, or joined to form macromolecules. This process often consists of metabolic pathways. Some of these biosynthetic pathways are located within a single cellular organelle, while others involve enzymes that are located within multiple cellular organelles. Examples of these biosynthetic pathways include the production of lipid membrane components and nucleotides. Biosynthesis is usually synonymous with anabolism.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.

Thiolases, also known as acetyl-coenzyme A acetyltransferases (ACAT), are enzymes which convert two units of acetyl-CoA to acetoacetyl CoA in the mevalonate pathway.

The transsulfuration pathway is a metabolic pathway involving the interconversion of cysteine and homocysteine through the intermediate cystathionine. Two transsulfurylation pathways are known: the forward and the reverse.

In enzymology, a homoserine dehydrogenase (EC 1.1.1.3) is an enzyme that catalyzes the chemical reaction
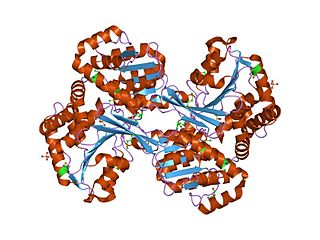
In enzymology, histidinol dehydrogenase (HIS4) (HDH) (EC 1.1.1.23) is an enzyme that catalyzes the chemical reaction

In enzymology, an aspartate-semialdehyde dehydrogenase is an enzyme that is very important in the biosynthesis of amino acids in prokaryotes, fungi, and some higher plants. It forms an early branch point in the metabolic pathway forming lysine, methionine, leucine and isoleucine from aspartate. This pathway also produces diaminopimelate which plays an essential role in bacterial cell wall formation. There is particular interest in ASADH as disabling this enzyme proves fatal to the organism giving rise to the possibility of a new class of antibiotics, fungicides, and herbicides aimed at inhibiting it.
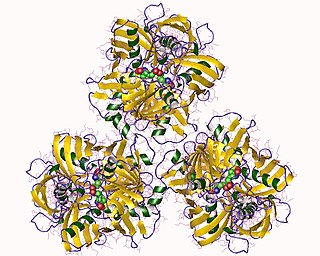
In enzymology, a diaminopimelate epimerase is an enzyme that catalyzes the chemical reaction
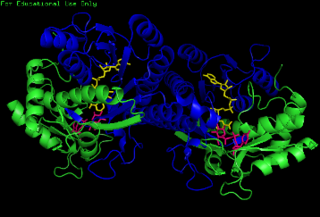
The enzyme UDP-glucose 4-epimerase, also known as UDP-galactose 4-epimerase or GALE, is a homodimeric epimerase found in bacterial, fungal, plant, and mammalian cells. This enzyme performs the final step in the Leloir pathway of galactose metabolism, catalyzing the reversible conversion of UDP-galactose to UDP-glucose. GALE tightly binds nicotinamide adenine dinucleotide (NAD+), a co-factor required for catalytic activity.

Isocitrate lyase, or ICL, is an enzyme in the glyoxylate cycle that catalyzes the cleavage of isocitrate to succinate and glyoxylate. Together with malate synthase, it bypasses the two decarboxylation steps of the tricarboxylic acid cycle and is used by bacteria, fungi, and plants.

In enzymology, an aminodeoxychorismate synthase is an enzyme that catalyzes the chemical reaction
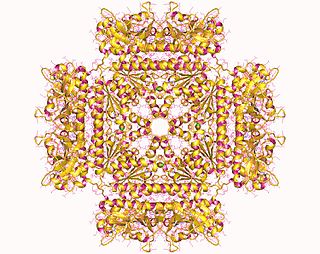

In molecular biology, the protein domain SAICAR synthase is an enzyme which catalyses a reaction to create SAICAR. In enzymology, this enzyme is also known as phosphoribosylaminoimidazolesuccinocarboxamide synthase. It is an enzyme that catalyzes the chemical reaction
In enzymology, a [acyl-carrier-protein] S-acetyltransferase is an enzyme that catalyzes the reversible chemical reaction

In enzymology, a malate synthase (EC 2.3.3.9) is an enzyme that catalyzes the chemical reaction
In enzymology, a maltose O-acetyltransferase is an enzyme that catalyzes the chemical reaction
In enzymology, a N6-hydroxylysine O-acetyltransferase (EC 2.3.1.102) is an enzyme that catalyzes the chemical reaction
In enzymology, a cysteine synthase is an enzyme that catalyzes the chemical reaction
In molecular biology, the Cys/Met metabolism PLP-dependent enzyme family is a family of proteins including enzymes involved in cysteine and methionine metabolism which use PLP (pyridoxal-5'-phosphate) as a cofactor.