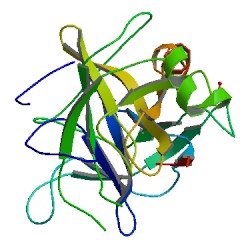
Chymotrypsin (EC 3.4.21.1, chymotrypsins A and B, alpha-chymar ophth, avazyme, chymar, chymotest, enzeon, quimar, quimotrase, alpha-chymar, alpha-chymotrypsin A, alpha-chymotrypsin) is a digestive enzyme component of pancreatic juice acting in the duodenum, where it performs proteolysis, the breakdown of proteins and polypeptides. Chymotrypsin preferentially cleaves peptide amide bonds where the side chain of the amino acid N-terminal to the scissile amide bond (the P1 position) is a large hydrophobic amino acid (tyrosine, tryptophan, and phenylalanine). These amino acids contain an aromatic ring in their side chain that fits into a hydrophobic pocket (the S1 position) of the enzyme. It is activated in the presence of trypsin. The hydrophobic and shape complementarity between the peptide substrate P1 side chain and the enzyme S1 binding cavity accounts for the substrate specificity of this enzyme. Chymotrypsin also hydrolyzes other amide bonds in peptides at slower rates, particularly those containing leucine at the P1 position.
Hydrolase is a class of enzymes that commonly perform as biochemical catalysts that use water to break a chemical bond, which typically results in dividing a larger molecule into smaller molecules. Some common examples of hydrolase enzymes are esterases including lipases, phosphatases, glycosidases, peptidases, and nucleosidases.
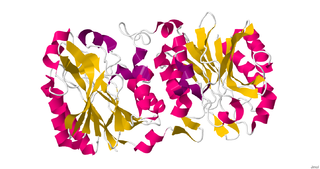
Nitrilase enzymes catalyse the hydrolysis of nitriles to carboxylic acids and ammonia, without the formation of "free" amide intermediates. Nitrilases are involved in natural product biosynthesis and post translational modifications in plants, animals, fungi and certain prokaryotes. Nitrilases can also be used as catalysts in preparative organic chemistry. Among others, nitrilases have been used for the resolution of racemic mixtures. Nitrilase should not be confused with nitrile hydratase which hydrolyses nitriles to amides. Nitrile hydratases are almost invariably co-expressed with an amidase, which converts the amide to the carboxylic acid. Consequently, it can sometimes be difficult to distinguish nitrilase activity from nitrile hydratase plus amidase activity.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.

Monoacylglycerol lipase is an enzyme that, in humans, is encoded by the MGLL gene. MAGL is a 33-kDa, membrane-associated member of the serine hydrolase superfamily and contains the classical GXSXG consensus sequence common to most serine hydrolases. The catalytic triad has been identified as Ser122, His269, and Asp239.
Serine hydrolases are one of the largest known enzyme classes comprising approximately ~200 enzymes or 1% of the genes in the human proteome. A defining characteristic of these enzymes is the presence of a particular serine at the active site, which is used for the hydrolysis of substrates. The hydrolysis of the ester or peptide bond proceeds in two steps. First, the acyl part of the substrate is transferred to the serine, making a new ester or amide bond and releasing the other part of the substrate is released. Later, in a slower step, the bond between the serine and the acyl group is hydrolyzed by water or hydroxide ion, regenerating free enzyme. Unlike other, non-catalytic, serines, the reactive serine of these hydrolases is typically activated by a proton relay involving a catalytic triad consisting of the serine, an acidic residue and a basic residue, although variations on this mechanism exist.
In enzymology, an acylagmatine amidase (EC 3.5.1.40) is an enzyme that catalyzes the chemical reaction
In enzymology, an aminoacylase (EC 3.5.1.14) is an enzyme that catalyzes the chemical reaction
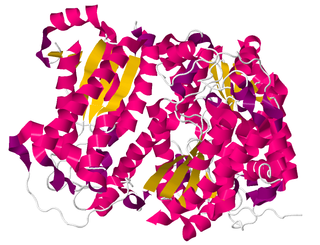
In enzymology, a creatininase (EC 3.5.2.10) is an enzyme that catalyses the hydrolysis of creatinine to creatine, which can then be metabolised to urea and sarcosine by creatinase.
In enzymology, a long-chain-fatty-acyl-glutamate deacylase (EC 3.5.1.55) is an enzyme that catalyzes the chemical reaction
In enzymology, an aculeacin-A deacylase is an enzyme that catalyzes the chemical reaction that cleaves the amide bond in aculeacin A and related neutral lipopeptide antibiotics, releasing the long-chain fatty acid side chain.

In enzymology, a peptide-N4-(N-acetyl-beta-glucosaminyl)asparagine amidase (EC 3.5.1.52) is an enzyme that catalyzes a chemical reaction that cleaves a N4-(acetyl-beta-D-glucosaminyl)asparagine residue in which the glucosamine residue may be further glycosylated, to yield a (substituted) N-acetyl-beta-D-glucosaminylamine and a peptide containing an aspartate residue. This enzyme belongs to the family of hydrolases, specifically those acting on carbon-nitrogen bonds other than peptide bonds in linear amides.
In enzymology, a N-formylglutamate deformylase (EC 3.5.1.68) is an enzyme that catalyzes the chemical reaction

In enzymology, a nicotinamidase (EC 3.5.1.19) is an enzyme that catalyzes the chemical reaction
In enzymology, a nicotinamide-nucleotide amidase (EC 3.5.1.42) is an enzyme that catalyzes the chemical reaction
In enzymology, an N-methylhydantoinase (ATP-hydrolysing) (EC 3.5.2.14) is an enzyme that catalyzes the chemical reaction

In enzymology, an omega-amidase (EC 3.5.1.3) is an enzyme that catalyzes the chemical reaction
In enzymology, a phthalyl amidase (EC 3.5.1.79) is an enzyme that catalyzes the chemical reaction
Dipeptidase E is an enzyme. This enzyme catalyses the following chemical reaction
N-acylethanolamine acid amide hydrolase (NAAA) EC 3.5.1.- is a member of the choloylglycine hydrolase family, a subset of the N-terminal nucleophile hydrolase superfamily. NAAA has a molecular weight of 31 kDa. The activation and inhibition of its catalytic site is of medical interest as a potential treatment for obesity and chronic pain. While it was discovered within the last decade, its structural similarity to the more familiar acid ceramidase (AC) and functional similarity to fatty acid amide hydrolase (FAAH) allow it to be studied extensively.









