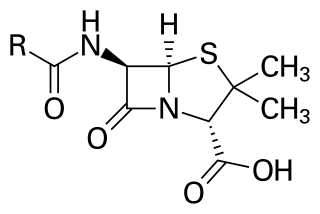
Penicillins are a group of β-lactam antibiotics originally obtained from Penicillium moulds, principally P. chrysogenum and P. rubens. Most penicillins in clinical use are synthesised by P. chrysogenum using deep tank fermentation and then purified. A number of natural penicillins have been discovered, but only two purified compounds are in clinical use: penicillin G and penicillin V. Penicillins were among the first medications to be effective against many bacterial infections caused by staphylococci and streptococci. They are still widely used today for different bacterial infections, though many types of bacteria have developed resistance following extensive use.

Staphylococcus aureus is a Gram-positive spherically shaped bacterium, a member of the Bacillota, and is a usual member of the microbiota of the body, frequently found in the upper respiratory tract and on the skin. It is often positive for catalase and nitrate reduction and is a facultative anaerobe that can grow without the need for oxygen. Although S. aureus usually acts as a commensal of the human microbiota, it can also become an opportunistic pathogen, being a common cause of skin infections including abscesses, respiratory infections such as sinusitis, and food poisoning. Pathogenic strains often promote infections by producing virulence factors such as potent protein toxins, and the expression of a cell-surface protein that binds and inactivates antibodies. S. aureus is one of the leading pathogens for deaths associated with antimicrobial resistance and the emergence of antibiotic-resistant strains, such as methicillin-resistant S. aureus (MRSA), is a worldwide problem in clinical medicine. Despite much research and development, no vaccine for S. aureus has been approved.

Methicillin-resistant Staphylococcus aureus (MRSA) is a group of gram-positive bacteria that are genetically distinct from other strains of Staphylococcus aureus. MRSA is responsible for several difficult-to-treat infections in humans. It caused more than 100,000 deaths worldwide attributable to antimicrobial resistance in 2019.

Methicillin (USAN), also known as meticillin (INN), is a narrow-spectrum β-lactam antibiotic of the penicillin class.

Pseudomonas fluorescens is a common Gram-negative, rod-shaped bacterium. It belongs to the Pseudomonas genus; 16S rRNA analysis as well as phylogenomic analysis has placed P. fluorescens in the P. fluorescens group within the genus, to which it lends its name.

Rifampicin, also known as rifampin, is an ansamycin antibiotic used to treat several types of bacterial infections, including tuberculosis (TB), Mycobacterium avium complex, leprosy, and Legionnaires' disease. It is almost always used together with other antibiotics with two notable exceptions: when given as a "preferred treatment that is strongly recommended" for latent TB infection; and when used as post-exposure prophylaxis to prevent Haemophilus influenzae type b and meningococcal disease in people who have been exposed to those bacteria. Before treating a person for a long period of time, measurements of liver enzymes and blood counts are recommended. Rifampicin may be given either by mouth or intravenously.

Fusidic acid, sold under the brand names Fucidin among others, is a steroid antibiotic that is often used topically in creams or ointments and eyedrops but may also be given systemically as tablets or injections.
As of October 2008, the global problem of advancing antimicrobial resistance has led to a renewed interest in its use.

Vancomycin-resistant Staphylococcus aureus (VRSA) are strains of Staphylococcus aureus that have acquired resistance to the glycopeptide antibiotic vancomycin. Bacteria can acquire resistant genes either by random mutation or through the transfer of DNA from one bacterium to another. Resistance genes interfere with the normal antibiotic function and allow a bacteria to grow in the presence of the antibiotic. Resistance in VRSA is conferred by the plasmid-mediated vanA gene and operon. Although VRSA infections are uncommon, VRSA is often resistant to other types of antibiotics and a potential threat to public health because treatment options are limited. VRSA is resistant to many of the standard drugs used to treat S. aureus infections. Furthermore, resistance can be transferred from one bacterium to another.
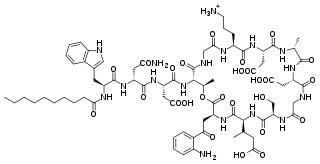
Daptomycin, sold under the brand name Cubicin among others, is a lipopeptide antibiotic used in the treatment of systemic and life-threatening infections caused by Gram-positive organisms.

Staphylococcus haemolyticus is a member of the coagulase-negative staphylococci (CoNS). It is part of the skin flora of humans, and its largest populations are usually found at the axillae, perineum, and inguinal areas. S. haemolyticus also colonizes primates and domestic animals. It is a well-known opportunistic pathogen, and is the second-most frequently isolated CoNS. Infections can be localized or systemic, and are often associated with the insertion of medical devices. The highly antibiotic-resistant phenotype and ability to form biofilms make S. haemolyticus a difficult pathogen to treat. Its most closely related species is Staphylococcus borealis.

Pseudomonas aeruginosa is a common encapsulated, Gram-negative, aerobic–facultatively anaerobic, rod-shaped bacterium that can cause disease in plants and animals, including humans. A species of considerable medical importance, P. aeruginosa is a multidrug resistant pathogen recognized for its ubiquity, its intrinsically advanced antibiotic resistance mechanisms, and its association with serious illnesses – hospital-acquired infections such as ventilator-associated pneumonia and various sepsis syndromes. P. aeruginosa is able to selectively inhibit various antibiotics from penetrating its outer membrane - and has high resistance to several antibiotics, according to the World Health Organization P. aeruginosa poses one of the greatest threats to humans in terms of antibiotic resistance.
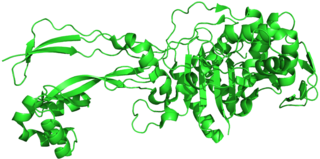
Penicillin-binding proteins (PBPs) are a group of proteins that are characterized by their affinity for and binding of penicillin. They are a normal constituent of many bacteria; the name just reflects the way by which the protein was discovered. All β-lactam antibiotics bind to PBPs, which are essential for bacterial cell wall synthesis. PBPs are members of a subgroup of enzymes called transpeptidases. Specifically, PBPs are DD-transpeptidases.
The rpoB gene encodes the β subunit of bacterial RNA polymerase and the homologous plastid-encoded RNA polymerase (PEP). It codes for 1342 amino acids in E. coli, making it the second-largest polypeptide in the bacterial cell. It is targeted by the rifamycin family of antibacterials, such as rifampin. Mutations in rpoB that confer resistance to rifamycins do so by altering the protein's drug-binding residues, thereby reducing affinity for these antibiotics.
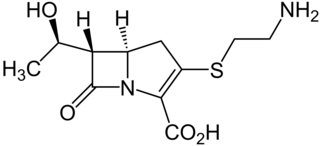
Thienamycin is one of the most potent naturally produced antibiotics known thus far, discovered in Streptomyces cattleya in 1976. Thienamycin has excellent activity against both Gram-positive and Gram-negative bacteria and is resistant to bacterial β-lactamase enzymes. Thienamycin is a zwitterion at pH 7.
mecA is a gene found in bacterial cells which allows them to be resistant to antibiotics such as methicillin, penicillin and other penicillin-like antibiotics.

Arbekacin (INN) is a semisynthetic aminoglycoside antibiotic which was derived from kanamycin. It is primarily used for the treatment of infections caused by multi-resistant bacteria including methicillin-resistant Staphylococcus aureus (MRSA). Arbekacin was originally synthesized from dibekacin in 1973 by Hamao Umezawa and collaborators. It has been registered and marketed in Japan since 1990 under the trade name Habekacin. Arbekacin is no longer covered by patent and generic versions of the drug are also available under such trade names as Decontasin and Blubatosine.
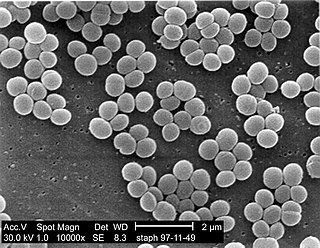
Staphylococcus is a genus of Gram-positive bacteria in the family Staphylococcaceae from the order Bacillales. Under the microscope, they appear spherical (cocci), and form in grape-like clusters. Staphylococcus species are facultative anaerobic organisms.
Streptogramin B is a subgroup of the streptogramin antibiotics family. These natural products are cyclic hexa- or hepta depsipeptides produced by various members of the genus of bacteria Streptomyces. Many of the members of the streptogramins reported in the literature have the same structure and different names; for example, pristinamycin IA = vernamycin Bα = mikamycin B = osteogrycin B.
Teixobactin is a peptide-like secondary metabolite of some species of bacteria, that kills some gram-positive bacteria. It appears to belong to a new class of antibiotics, and harms bacteria by binding to lipid II and lipid III, important precursor molecules for forming the cell wall.
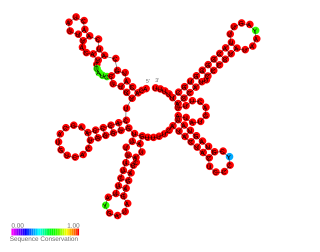
In molecular biology the small pathogenicity island RNA X gene is a bacterial non-coding RNA. It was discovered in a large-scale analysis of Staphylococcus aureus. SprX was shown to influence antibiotic resistance of the bacteria to Vancomycin and Teicoplanin glycopeptides, which are used to treat MRSA infections. In this study the authors identified a SprX target, stage V sporulation protein G. By reducing Spo VG expression levels, SprX affects S. aureus resistance to the glycopeptide antibiotics. Further work demonstrated its involvement in the regulation of pathogenicity factors.























