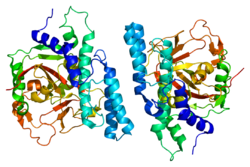
Breast cancer type 1 susceptibility protein is a protein that in humans is encoded by the BRCA1 gene. Orthologs are common in other vertebrate species, whereas invertebrate genomes may encode a more distantly related gene. BRCA1 is a human tumor suppressor gene and is responsible for repairing DNA.

DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encodes its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA damage, resulting in tens of thousands of individual molecular lesions per cell per day. Many of these lesions cause structural damage to the DNA molecule and can alter or eliminate the cell's ability to transcribe the gene that the affected DNA encodes. Other lesions induce potentially harmful mutations in the cell's genome, which affect the survival of its daughter cells after it undergoes mitosis. As a consequence, the DNA repair process is constantly active as it responds to damage in the DNA structure. When normal repair processes fail, and when cellular apoptosis does not occur, irreparable DNA damage may occur, including double-strand breaks and DNA crosslinkages. This can eventually lead to malignant tumors, or cancer as per the two-hit hypothesis.

Poly (ADP-ribose) polymerase (PARP) is a family of proteins involved in a number of cellular processes such as DNA repair, genomic stability, and programmed cell death.
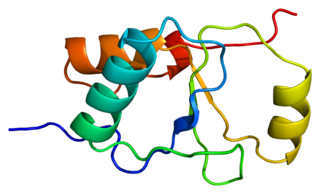
DNA repair protein XRCC1, also known as X-ray repair cross-complementing protein 1, is a protein that in humans is encoded by the XRCC1 gene. XRCC1 is involved in DNA repair, where it complexes with DNA ligase III.

Histone-modifying enzymes are enzymes involved in the modification of histone substrates after protein translation and affect cellular processes including gene expression. To safely store the eukaryotic genome, DNA is wrapped around four core histone proteins, which then join to form nucleosomes. These nucleosomes further fold together into highly condensed chromatin, which renders the organism's genetic material far less accessible to the factors required for gene transcription, DNA replication, recombination and repair. Subsequently, eukaryotic organisms have developed intricate mechanisms to overcome this repressive barrier imposed by the chromatin through histone modification, a type of post-translational modification which typically involves covalently attaching certain groups to histone residues. Once added to the histone, these groups elicit either a loose and open histone conformation, euchromatin, or a tight and closed histone conformation, heterochromatin. Euchromatin marks active transcription and gene expression, as the light packing of histones in this way allows entry for proteins involved in the transcription process. As such, the tightly packed heterochromatin marks the absence of current gene expression.

ADP-ribosylation is the addition of one or more ADP-ribose moieties to a protein. It is a reversible post-translational modification that is involved in many cellular processes, including cell signaling, DNA repair, gene regulation and apoptosis. Improper ADP-ribosylation has been implicated in some forms of cancer. It is also the basis for the toxicity of bacterial compounds such as cholera toxin, diphtheria toxin, and others.

Protein C-ets-1 is a protein that in humans is encoded by the ETS1 gene. The protein encoded by this gene belongs to the ETS family of transcription factors.
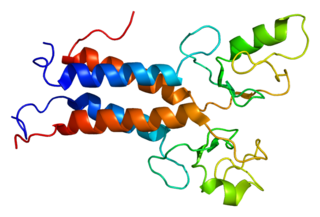
BRCA1-associated RING domain protein 1 is a protein that in humans is encoded by the BARD1 gene. The human BARD1 protein is 777 amino acids long and contains a RING finger domain, four ankyrin repeats, and two tandem BRCT domains.
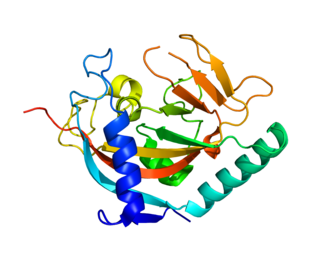
Tankyrase, also known as tankyrase 1, is an enzyme that in humans is encoded by the TNKS gene. It inhibits the binding of TERF1 to telomeric DNA. Tankyrase attracts substantial interest in cancer research through its interaction with AXIN1 and AXIN2, which are negative regulators of pro-oncogenic β-catenin signaling. Importantly, activity in the β-catenin destruction complex can be increased by tankyrase inhibitors and thus such inhibitors are a potential therapeutic option to reduce the growth of β-catenin-dependent cancers.

Poly [ADP-ribose] polymerase 4 is an enzyme that in humans is encoded by the PARP4 gene.
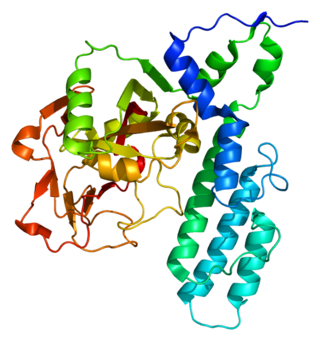
Poly [ADP-ribose] polymerase 3 is an enzyme that in humans is encoded by the PARP3 gene.

Poly [ADP-ribose] polymerase 2 is an enzyme that in humans is encoded by the PARP2 gene. It is one of the PARP family of enzymes.
Post-transcriptional regulation is the control of gene expression at the RNA level. It occurs once the RNA polymerase has been attached to the gene's promoter and is synthesizing the nucleotide sequence. Therefore, as the name indicates, it occurs between the transcription phase and the translation phase of gene expression. These controls are critical for the regulation of many genes across human tissues. It also plays a big role in cell physiology, being implicated in pathologies such as cancer and neurodegenerative diseases.

Olaparib, sold under the brand name Lynparza, is a medication for the maintenance treatment of BRCA-mutated advanced ovarian cancer in adults. It is a PARP inhibitor, inhibiting poly ADP ribose polymerase (PARP), an enzyme involved in DNA repair. It acts against cancers in people with hereditary BRCA1 or BRCA2 mutations, which include some ovarian, breast, and prostate cancers.

PARP inhibitors are a group of pharmacological inhibitors of the enzyme poly ADP ribose polymerase (PARP).

3-Aminobenzamide is a benzamide. It is an off-white powder and has the chemical formula C7H8N2O.
Parthanatos is a form of programmed cell death that is distinct from other cell death processes such as necrosis and apoptosis. While necrosis is caused by acute cell injury resulting in traumatic cell death and apoptosis is a highly controlled process signalled by apoptotic intracellular signals, parthanatos is caused by the accumulation of Poly(ADP ribose) (PAR) and the nuclear translocation of apoptosis-inducing factor (AIF) from mitochondria. Parthanatos is also known as PARP-1 dependent cell death. PARP-1 mediates parthanatos when it is over-activated in response to extreme genomic stress and synthesizes PAR which causes nuclear translocation of AIF. Parthanatos is involved in diseases that afflict hundreds of millions of people worldwide. Well known diseases involving parthanatos include Parkinson's disease, stroke, heart attack, and diabetes. It also has potential use as a treatment for ameliorating disease and various medical conditions such as diabetes and obesity.

Talazoparib, sold under the brand name Talzenna, is an orally available poly ADP ribose polymerase (PARP) inhibitor marketed by Pfizer for the treatment of advanced breast cancer with germline BRCA mutations. Talazoparib is similar to the first in class PARP inhibitor, olaparib. It was approved in October 2018, in the United States and June 2019, in the European Union for germline BRCA-mutated, HER2-negative locally advanced or metastatic breast cancer.

7-Methylguanine is a modified purine nucleobase. It is a methylated version of guanine. The 7-methylguanine nucleoside is called 7-methylguanosine. However, the free 7-methylguanine base is not involved in the synthesis of nucleotides and not incorporated directly into nucleic acids. 7-Methylguanine is a natural inhibitor of poly (ADP-ribose) polymerase (PARP) and tRNA guanine transglycosylase (TGT) - and thus may exert anticancer activity. For example, it was demonstrated that 7-methylguanine could accelerate apoptotic death of BRCA1-deficient breast cancer cells induced by cisplatin and doxorubicin.

HRDetect is a whole-genome sequencing (WGS)-based classifier designed to predict BRCA1 and BRCA2 deficiency based on six mutational signatures. Additionally, the classifier is able to identify similarities in mutational profiles of tumors to that of tumors with BRCA1 and BRCA2 defects, also known as BRCAness. This classifier can be applied to assess the implementation of PARP inhibitors in patients with BRCA1/BRCA2 deficiency. The final output is a probability of BRCA1/2 mutation.