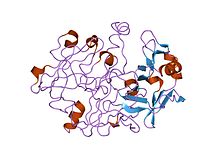
A protease is an enzyme that catalyzes proteolysis, the breakdown of proteins into smaller polypeptides or single amino acids. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks bonds. Proteases are involved in many biological functions, including digestion of ingested proteins, protein catabolism, and cell signaling.
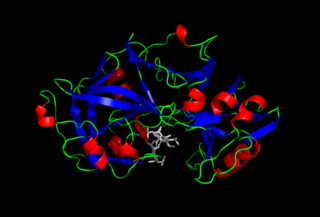
Pepsin is an endopeptidase that breaks down proteins into smaller peptides. It is produced in the chief cells of the stomach lining and is one of the main digestive enzymes in the digestive systems of humans and many other animals, where it helps digest the proteins in food. Pepsin is an aspartic protease, using a catalytic aspartate in its active site.
In biology and biochemistry, protease inhibitors, or antiproteases, are molecules that inhibit the function of proteases. Many naturally occurring protease inhibitors are proteins.
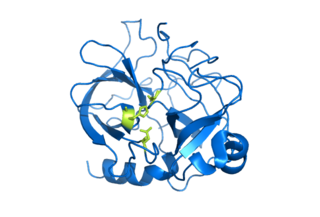
Serine proteases are enzymes that cleave peptide bonds in proteins, in which serine serves as the nucleophilic amino acid at the (enzyme's) active site. They are found ubiquitously in both eukaryotes and prokaryotes. Serine proteases fall into two broad categories based on their structure: chymotrypsin-like (trypsin-like) or subtilisin-like.
A metalloproteinase, or metalloprotease, is any protease enzyme whose catalytic mechanism involves a metal. An example of this would be ADAM12 which plays a significant role in the fusion of muscle cells during embryo development, in a process known as myogenesis.

Papain, also known as papaya proteinase I, is a cysteine protease enzyme present in papaya and mountain papaya.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An Acid-Base-Nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.
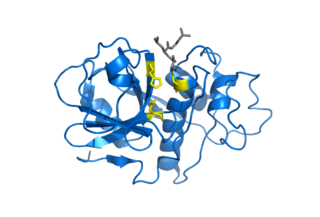
Cysteine proteases, also known as thiol proteases, are enzymes that degrade proteins. These proteases share a common catalytic mechanism that involves a nucleophilic cysteine thiol in a catalytic triad or dyad.

Beta-secretase 1, also known as beta-site amyloid precursor protein cleaving enzyme 1, beta-site APP cleaving enzyme 1 (BACE1), membrane-associated aspartic protease 2, memapsin-2, aspartyl protease 2, and ASP2, is an enzyme that in humans is encoded by the BACE1 gene. Expression of BACE1 is observed mainly in neurons.

Pepsin A is an enzyme. This enzyme catalyses the following chemical reaction
In molecular biology, the Signal Peptide Peptidase (SPP) is a type of protein that specifically cleaves parts of other proteins. It is an intramembrane aspartyl protease with the conserved active site motifs 'YD' and 'GxGD' in adjacent transmembrane domains (TMDs). Its sequences is highly conserved in different vertebrate species. SPP cleaves remnant signal peptides left behind in membrane by the action of signal peptidase and also plays key roles in immune surveillance and the maturation of certain viral proteins.

Cathepsin D is a protein that in humans is encoded by the CTSD gene. This gene encodes a lysosomal aspartyl protease composed of a protein dimer of disulfide-linked heavy and light chains, both produced from a single protein precursor. Cathepsin D is an aspartic endo-protease that is ubiquitously distributed in lysosomes. The main function of cathepsin D is to degrade proteins and activate precursors of bioactive proteins in pre-lysosomal compartments. This proteinase, which is a member of the peptidase A1 family, has a specificity similar to but narrower than that of pepsin A. Transcription of the CTSD gene is initiated from several sites, including one that is a start site for an estrogen-regulated transcript. Mutations in this gene are involved in the pathogenesis of several diseases, including breast cancer and possibly Alzheimer disease. Homozygous deletion of the CTSD gene leads to early lethality in the postnatal phase. Deficiency of CTSD gene has been reported an underlying cause of neuronal ceroid lipofuscinosis (NCL).

Cathepsin E is an enzyme that in humans is encoded by the CTSE gene. The enzyme is also known as slow-moving proteinase, erythrocyte membrane aspartic proteinase, SMP, EMAP, non-pepsin proteinase, cathepsin D-like acid proteinase, cathepsin E-like acid proteinase, cathepsin D-type proteinase) is an enzyme.

Cathepsin F is a protein that in humans is encoded by the CTSF gene.
Nepenthesin is an aspartic protease of plant origin that has so far been identified in the pitcher secretions of Nepenthes and in the leaves of Drosera peltata. It is similar to pepsin, but differs in that it also cleaves on either side of Asp residues and at Lys┼Arg. While more pH and temperature stable than porcine pepsin A, it is considerably less stable in urea or guanidine hydrochloride. It is the only known protein with such a stability profile.
Retroviral aspartyl proteases or retropepsins are single domain aspartyl proteases from retroviruses, retrotransposons, and badnaviruses. These proteases are generally part of a larger pol or gag polyprotein. Retroviral proteases are homologous to a single domain of the two-domain eukaryotic aspartyl proteases such as pepsins, cathepsins, and renins. Retropepsins are members of MEROPS A2, clan AA. All known members are endopeptidases.
Rhizopuspepsin is an enzyme. This enzyme catalyses the following chemical reaction

Scytalidocarboxyl peptidase B, also known as Scytalidoglutamic peptidase and Scytalidopepsin B is a proteolytic enzyme. It was previously thought to be an aspartic protease, but determination of its molecular structure showed it to belong a novel group of proteases, glutamic protease.

Glutamic proteases are a group of proteolytic enzymes containing a glutamic acid residue within the active site. This type of protease was first described in 2004 and became the sixth catalytic type of protease. Members of this group of protease had been previously assumed to be an aspartate protease, but structural determination showed it to belong to a novel protease family. The first structure of this group of protease was scytalidoglutamic peptidase, the active site of which contains a catalytic dyad, glutamic acid (E) and glutamine (Q), which give rise to the name eqolisin. This group of proteases are found primarily in pathogenic fungi affecting plant and human.
Asparagine peptide lyase are one of the seven groups in which proteases, also termed proteolytic enzymes, peptidases, or proteinases, are classified according to their catalytic residue. The catalytic mechanism of the asparagine peptide lyases involves an asparagine residue acting as nucleophile to perform a nucleophilic elimination reaction, rather than hydrolysis, to catalyse the breaking of a peptide bond.