Related Research Articles
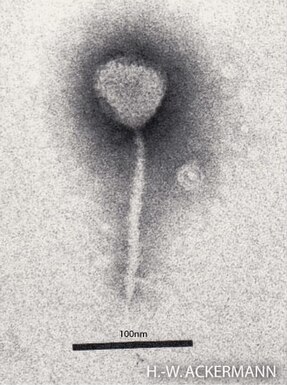
Enterobacteria phage λ is a bacterial virus, or bacteriophage, that infects the bacterial species Escherichia coli. It was discovered by Esther Lederberg in 1950. The wild type of this virus has a temperate life cycle that allows it to either reside within the genome of its host through lysogeny or enter into a lytic phase, during which it kills and lyses the cell to produce offspring. Lambda strains, mutated at specific sites, are unable to lysogenize cells; instead, they grow and enter the lytic cycle after superinfecting an already lysogenized cell.
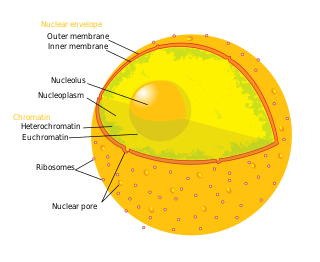
The nucleolus is the largest structure in the nucleus of eukaryotic cells. It is best known as the site of ribosome biogenesis. Nucleoli also participate in the formation of signal recognition particles and play a role in the cell's response to stress. Nucleoli are made of proteins, DNA and RNA and form around specific chromosomal regions called nucleolar organizing regions. Malfunction of nucleoli can be the cause of several human conditions called "nucleolopathies" and the nucleolus is being investigated as a target for cancer chemotherapy.

Transcription is the process of copying a segment of DNA into RNA. The segments of DNA transcribed into RNA molecules that can encode proteins are said to produce messenger RNA (mRNA). Other segments of DNA are copied into RNA molecules called non-coding RNAs (ncRNAs). Averaged over multiple cell types in a given tissue, the quantity of mRNA is more than 10 times the quantity of ncRNA. The general preponderance of mRNA in cells is valid even though less than 2% of the human genome can be transcribed into mRNA, while at least 80% of mammalian genomic DNA can be actively transcribed, with the majority of this 80% considered to be ncRNA.

In molecular biology, RNA polymerase, is an enzyme that synthesizes RNA from a DNA template.
In molecular biology and genetics, transcriptional regulation is the means by which a cell regulates the conversion of DNA to RNA (transcription), thereby orchestrating gene activity. A single gene can be regulated in a range of ways, from altering the number of copies of RNA that are transcribed, to the temporal control of when the gene is transcribed. This control allows the cell or organism to respond to a variety of intra- and extracellular signals and thus mount a response. Some examples of this include producing the mRNA that encode enzymes to adapt to a change in a food source, producing the gene products involved in cell cycle specific activities, and producing the gene products responsible for cellular differentiation in multicellular eukaryotes, as studied in evolutionary developmental biology.
A sigma factor is a protein needed for initiation of transcription in bacteria. It is a bacterial transcription initiation factor that enables specific binding of RNA polymerase (RNAP) to gene promoters. It is homologous to archaeal transcription factor B and to eukaryotic factor TFIIB. The specific sigma factor used to initiate transcription of a given gene will vary, depending on the gene and on the environmental signals needed to initiate transcription of that gene. Selection of promoters by RNA polymerase is dependent on the sigma factor that associates with it. They are also found in plant chloroplasts as a part of the bacteria-like plastid-encoded polymerase (PEP).
In genetics, a transcription terminator is a section of nucleic acid sequence that marks the end of a gene or operon in genomic DNA during transcription. This sequence mediates transcriptional termination by providing signals in the newly synthesized transcript RNA that trigger processes which release the transcript RNA from the transcriptional complex. These processes include the direct interaction of the mRNA secondary structure with the complex and/or the indirect activities of recruited termination factors. Release of the transcriptional complex frees RNA polymerase and related transcriptional machinery to begin transcription of new mRNAs.

Ribosomal DNA (rDNA) is a DNA sequence that codes for ribosomal RNA. These sequences regulate transcription initiation and amplification, and contain both transcribed and non-transcribed spacer segments.

The preinitiation complex is a complex of approximately 100 proteins that is necessary for the transcription of protein-coding genes in eukaryotes and archaea. The preinitiation complex positions RNA polymerase II at gene transcription start sites, denatures the DNA, and positions the DNA in the RNA polymerase II active site for transcription.

RNA polymerase II is a multiprotein complex that transcribes DNA into precursors of messenger RNA (mRNA) and most small nuclear RNA (snRNA) and microRNA. It is one of the three RNAP enzymes found in the nucleus of eukaryotic cells. A 550 kDa complex of 12 subunits, RNAP II is the most studied type of RNA polymerase. A wide range of transcription factors are required for it to bind to upstream gene promoters and begin transcription.
General transcription factors (GTFs), also known as basal transcriptional factors, are a class of protein transcription factors that bind to specific sites (promoter) on DNA to activate transcription of genetic information from DNA to messenger RNA. GTFs, RNA polymerase, and the mediator constitute the basic transcriptional apparatus that first bind to the promoter, then start transcription. GTFs are also intimately involved in the process of gene regulation, and most are required for life.
In eukaryote cells, RNA polymerase III transcribes DNA to synthesize ribosomal 5S rRNA, tRNA and other small RNAs.

Ribosome biogenesis is the process of making ribosomes. In prokaryotes, this process takes place in the cytoplasm with the transcription of many ribosome gene operons. In eukaryotes, it takes place both in the cytoplasm and in the nucleolus. It involves the coordinated function of over 200 proteins in the synthesis and processing of the three prokaryotic or four eukaryotic rRNAs, as well as assembly of those rRNAs with the ribosomal proteins. Most of the ribosomal proteins fall into various energy-consuming enzyme families including ATP-dependent RNA helicases, AAA-ATPases, GTPases, and kinases. About 60% of a cell's energy is spent on ribosome production and maintenance.
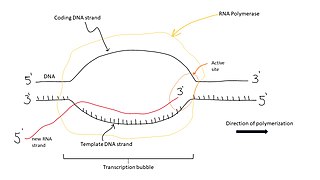
A transcription bubble is a molecular structure formed during DNA transcription when a limited portion of the DNA double strand is unwound. The size of a transcription bubble ranges from 12-14 base pairs. A transcription bubble is formed when the RNA polymerase enzyme binds to a promoter and causes two DNA strands to detach. It presents a region of unpaired DNA, where a short stretch of nucleotides are exposed on each strand of the double helix.

Bacterial transcription is the process in which a segment of bacterial DNA is copied into a newly synthesized strand of messenger RNA (mRNA) with use of the enzyme RNA polymerase. The process occurs in three main steps: initiation, elongation, and termination; and the end result is a strand of mRNA that is complementary to a single strand of DNA. Generally, the transcribed region accounts for more than one gene. In fact, many prokaryotic genes occur in operons, which are a series of genes that work together to code for the same protein or gene product and are controlled by a single promoter. Bacterial RNA polymerase is made up of four subunits and when a fifth subunit attaches, called the σ-factor, the polymerase can recognize specific binding sequences in the DNA, called promoters. The binding of the σ-factor to the promoter is the first step in initiation. Once the σ-factor releases from the polymerase, elongation proceeds. The polymerase continues down the double stranded DNA, unwinding it and synthesizing the new mRNA strand until it reaches a termination site. There are two termination mechanisms that are discussed in further detail below. Termination is required at specific sites for proper gene expression to occur. Gene expression determines how much gene product, such as protein, is made by the gene. Transcription is carried out by RNA polymerase but its specificity is controlled by sequence-specific DNA binding proteins called transcription factors. Transcription factors work to recognize specific DNA sequences and based on the cells needs, promote or inhibit additional transcription.

Eukaryotic transcription is the elaborate process that eukaryotic cells use to copy genetic information stored in DNA into units of transportable complementary RNA replica. Gene transcription occurs in both eukaryotic and prokaryotic cells. Unlike prokaryotic RNA polymerase that initiates the transcription of all different types of RNA, RNA polymerase in eukaryotes comes in three variations, each translating a different type of gene. A eukaryotic cell has a nucleus that separates the processes of transcription and translation. Eukaryotic transcription occurs within the nucleus where DNA is packaged into nucleosomes and higher order chromatin structures. The complexity of the eukaryotic genome necessitates a great variety and complexity of gene expression control.

Nucleolar transcription factor 1 is a protein that in humans is encoded by the UBTF gene.

TATA box-binding protein-associated factor RNA polymerase I subunit C is an enzyme that in humans is encoded by the TAF1C gene.
RNA polymerase II holoenzyme is a form of eukaryotic RNA polymerase II that is recruited to the promoters of protein-coding genes in living cells. It consists of RNA polymerase II, a subset of general transcription factors, and regulatory proteins known as SRB proteins.

Selective factor 1 is a transcription factor that binds to the promoter of genes and recruits a preinitiation complex to which RNA polymerase I will bind to and begin the transcription of ribosomal RNA (rRNA).
References
- ↑ Russell, Jackie; Zomerdijk, Joost C B M (2006). "The RNA polymerase I transcription machinery". Biochemical Society Symposium. 73 (73): 203–16. doi:10.1042/bss0730203. PMC 3858827 . PMID 16626300.
- ↑ Engel, Christoph; Sainsbury, Sarah; Cheung, Alan C.; Kostrewa, Dirk; Cramer, Patrick (23 October 2013). "RNA polymerase I structure and transcription regulation". Nature. 502 (7473): 650–655. Bibcode:2013Natur.502..650E. doi:10.1038/nature12712. hdl: 11858/00-001M-0000-0015-3B48-5 . PMID 24153182. S2CID 205236187.
- ↑ Zentner, Gabriel E; Saiakhova, Alina; Manaenkov, Pavel; Adams, Mark D; Scacheri, Peter C (25 February 2011). "Integrative genomic analysis of human ribosomal DNA". Nucleic Acids Research. 39 (12): 4949–4960. doi:10.1093/nar/gkq1326. PMC 3130253 . PMID 21355038.
- ↑ Edger, Patrick P; Tang, Michelle; Bird, Kevin A; Mayfield, Dustin R; Conant, Gavin; Mummenhoff, Klaus; Koch, Marcus A; Pires, J Chris (1 July 2014). "Secondary structure analyses of the nuclear rRNA internal transcribed spacers and assessment of its phylogenetic utility across the Brassicaceae (mustards)". PLOS ONE. 9 (7): e101341. Bibcode:2014PLoSO...9j1341E. doi: 10.1371/journal.pone.0101341 . PMC 4077792 . PMID 24984034.
- ↑ Appling, Dean; Anthony-Cahill, Spencer; Mathews, Christopher (2016). Biochemistry: Concepts and Connections. Hoboken, New Jersey: Pearson. p. 742. ISBN 978-0-321-83992-3.
- ↑ Watkins, Nicholas J.; Bohnsack, Markus T. (May 2012). "The box C/D and H/ACA snoRNPs: key players in the modification, processing and the dynamic folding of ribosomal RNA". Wiley Interdisciplinary Reviews: RNA. 3 (3): 397–414. doi:10.1002/wrna.117. PMID 22065625. S2CID 25766812.
- 1 2 Venema, Jaap; Tollervey, David (December 1999). "Ribosome Synthesis in Saccharomyces cerevisiae". Annual Review of Genetics. 33 (1): 261–311. doi:10.1146/annurev.genet.33.1.261. PMID 10690410.
- ↑ Grandori, Carla; Gomez-Roman, Natividad; Felton-Edkins, Zoe A.; Ngouenet, Celine; Galloway, Denise A.; Eisenman, Robert N.; White, Robert J. (20 February 2005). "c-Myc binds to human ribosomal DNA and stimulates transcription of rRNA genes by RNA polymerase I". Nature Cell Biology. 7 (3): 311–318. doi:10.1038/ncb1224. PMID 15723054. S2CID 8913931.
- ↑ Jantzen, Hans-Michael; Admon, Arie; Bell, Stephen P.; Tjian, Robert (26 April 1990). "Nucleolar transcription factor hUBF contains a DNA-binding motif with homology to HMG proteins". Nature. 344 (6269): 830–836. Bibcode:1990Natur.344..830J. doi:10.1038/344830a0. PMID 2330041. S2CID 4280039.
- 1 2 Grummt, Ingrid (15 July 2003). "Life on a planet of its own: regulation of RNA polymerase I transcription in the nucleolus". Genes & Development. 17 (14): 1691–1702. doi: 10.1101/gad.1098503R . PMID 12865296 . Retrieved 16 December 2014.
- ↑ Learned, R Marc; Cordes, Sabine; Tjian, Robert (June 1985). "Purification and characterization of a transcription factor that confers promoter specificity to human RNA polymerase I". Molecular and Cellular Biology. 5 (6): 1358–69. doi:10.1128/MCB.5.6.1358. PMC 366865 . PMID 3929071.
- ↑ Clos, Joachim; Buttgereit, Detlev; Grummt, Ingrid (February 1986). "A purified transcription factor (TIF-IB) binds to essential sequences of the mouse rDNA promoter". Proceedings of the National Academy of Sciences of the United States of America. 83 (3): 604–8. Bibcode:1986PNAS...83..604C. doi: 10.1073/pnas.83.3.604 . PMC 322912 . PMID 3456157.
- ↑ Serizawa N, Horiuchi T, Kobayashi T (2004). "Transcription-mediated hyper-recombination in HOT1". Genes Cells. 9 (4): 305–15. doi: 10.1111/j.1356-9597.2004.00729.x . PMID 15066122.