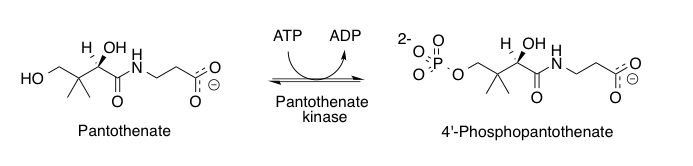
Pantothenic acid (vitamin B5) is a B vitamin and an essential nutrient. All animals need pantothenic acid in order to synthesize coenzyme A (CoA)—essential for metabolizing fatty acid—and to synthesize and metabolize proteins, carbohydrates, and fats.

Coenzyme A (CoA, SHCoA, CoASH) is a coenzyme, notable for its role in the synthesis and oxidation of fatty acids, and the oxidation of pyruvate in the citric acid cycle. All genomes sequenced to date encode enzymes that use coenzyme A as a substrate, and around 4% of cellular enzymes use it (or a thioester) as a substrate. In humans, CoA biosynthesis requires cysteine, pantothenate (vitamin B5), and adenosine triphosphate (ATP).

Carnitine is a quaternary ammonium compound involved in metabolism in most mammals, plants, and some bacteria. In support of energy metabolism, carnitine transports long-chain fatty acids from the cytosol into mitochondria to be oxidized for free energy production, and also participates in removing products of metabolism from cells. Given its key metabolic roles, carnitine is concentrated in tissues like skeletal and cardiac muscle that metabolize fatty acids as an energy source. Generally individuals, including strict vegetarians, synthesize enough L-carnitine in vivo.

Acetyl-CoA carboxylase (ACC) is a biotin-dependent enzyme that catalyzes the irreversible carboxylation of acetyl-CoA to produce malonyl-CoA through its two catalytic activities, biotin carboxylase (BC) and carboxyltransferase (CT). ACC is a multi-subunit enzyme in most prokaryotes and in the chloroplasts of most plants and algae, whereas it is a large, multi-domain enzyme in the cytoplasm of most eukaryotes. The most important function of ACC is to provide the malonyl-CoA substrate for the biosynthesis of fatty acids. The activity of ACC can be controlled at the transcriptional level as well as by small molecule modulators and covalent modification. The human genome contains the genes for two different ACCs—ACACA and ACACB.

Glucose-6-phosphate dehydrogenase (G6PD or G6PDH) (EC 1.1.1.49) is a cytosolic enzyme that catalyzes the chemical reaction
Pantothenate kinase-associated neurodegeneration (PKAN), formerly called Hallervorden–Spatz syndrome, is a genetic degenerative disease of the brain that can lead to parkinsonism, dystonia, dementia, and ultimately death. Neurodegeneration in PKAN is accompanied by an excess of iron that progressively builds up in the brain.
Fatty acid degradation is the process in which fatty acids are broken down into their metabolites, in the end generating acetyl-CoA, the entry molecule for the citric acid cycle, the main energy supply of living organisms, including bacteria and animals. It includes three major steps:
The long chain fatty acyl-CoA ligase is an enzyme of the ligase family that activates the oxidation of complex fatty acids. Long chain fatty acyl-CoA synthetase catalyzes the formation of fatty acyl-CoA by a two-step process proceeding through an adenylated intermediate. The enzyme catalyzes the following reaction,

Carnitine palmitoyltransferase I (CPT1) also known as carnitine acyltransferase I, CPTI, CAT1, CoA:carnitine acyl transferase (CCAT), or palmitoylCoA transferase I, is a mitochondrial enzyme responsible for the formation of acyl carnitines by catalyzing the transfer of the acyl group of a long-chain fatty acyl-CoA from coenzyme A to l-carnitine. The product is often Palmitoylcarnitine, but other fatty acids may also be substrates. It is part of a family of enzymes called carnitine acyltransferases. This "preparation" allows for subsequent movement of the acyl carnitine from the cytosol into the intermembrane space of mitochondria.

Acetyl-CoA acetyltransferase, mitochondrial, also known as acetoacetyl-CoA thiolase, is an enzyme that in humans is encoded by the ACAT1 gene.
ATP citrate synthase (also ATP citrate lyase (ACLY)) is an enzyme that in animals represents an important step in fatty acid biosynthesis. By converting citrate to acetyl-CoA, the enzyme links carbohydrate metabolism, which yields citrate as an intermediate, with fatty acid biosynthesis, which consumes acetyl-CoA. In plants, ATP citrate lyase generates cytosolic acetyl-CoA precursors of thousands of specialized metabolites, including waxes, sterols, and polyketides.

Carnitine O-acetyltransferase also called carnitine acetyltransferase is an enzyme that encoded by the CRAT gene that catalyzes the chemical reaction

Carnitine O-octanoyltransferase is a member of the transferase family, more specifically a carnitine acyltransferase, a type of enzyme which catalyzes the transfer of acyl groups from acyl-CoAs to carnitine, generating CoA and an acyl-carnitine. The systematic name of this enzyme is octanoyl-CoA:L-carnitine O-octanoyltransferase. Other names in common use include medium-chain/long-chain carnitine acyltransferase, carnitine medium-chain acyltransferase, easily solubilized mitochondrial carnitine palmitoyltransferase, and overt mitochondrial carnitine palmitoyltransferase. Specifically, CROT catalyzes the chemical reaction:

Pantothenate kinase 2, mitochondrial is an enzyme that in humans is encoded by the PANK2 gene.

Acyl-coenzyme A thioesterase 11 also known as StAR-related lipid transfer protein 14 (STARD14) is an enzyme that in humans is encoded by the ACOT11 gene. This gene encodes a protein with acyl-CoA thioesterase activity towards medium (C12) and long-chain (C18) fatty acyl-CoA substrates which relies on its StAR-related lipid transfer domain. Expression of a similar murine protein in brown adipose tissue is induced by cold exposure and repressed by warmth. Expression of the mouse protein has been associated with obesity, with higher expression found in obesity-resistant mice compared with obesity-prone mice. Alternative splicing results in two transcript variants encoding different isoforms.

Pantothenate kinase 4 is an enzyme that in humans is encoded by the PANK4 gene.

Pantothenate kinase 1 is an enzyme that in humans is encoded by the PANK1 gene.

Acyl-CoA thioesterase 1 is a protein that in humans is encoded by the ACOT1 gene.

The bump-and-hole method is a tool in chemical genetics for studying a specific isoform in a protein family without perturbing the other members of the family. The unattainability of isoform-selective inhibition due to structural homology in protein families is a major challenge of chemical genetics. With the bump-and-hole approach, a protein–ligand interface is engineered to achieve selectivity through steric complementarity while maintaining biochemical competence and orthogonality to the wild type pair. Typically, a "bumped" ligand/inhibitor analog is designed to bind a corresponding "hole-modified" protein. Bumped ligands are commonly bulkier derivatives of a cofactor of the target protein. Hole-modified proteins are recombinantly expressed with an amino acid substitution from a larger to smaller residue, e.g. glycine or alanine, at the cofactor binding site. The designed ligand/inhibitor has specificity for the engineered protein due to steric complementarity, but not the native counterpart due to steric interference.
Andrimid is an antibiotic natural product that is produced by the marine bacterium Vibrio coralliilyticus. Andrimid is an inhibitor of fatty acid biosynthesis by blocking the carboxyl transfer reaction of acetyl-CoA carboxylase (ACC).