
A DNA virus is a virus that has a genome made of deoxyribonucleic acid (DNA) that is replicated by a DNA polymerase. They can be divided between those that have two strands of DNA in their genome, called double-stranded DNA (dsDNA) viruses, and those that have one strand of DNA in their genome, called single-stranded DNA (ssDNA) viruses. dsDNA viruses primarily belong to two realms: Duplodnaviria and Varidnaviria, and ssDNA viruses are almost exclusively assigned to the realm Monodnaviria, which also includes some dsDNA viruses. Additionally, many DNA viruses are unassigned to higher taxa. Reverse transcribing viruses, which have a DNA genome that is replicated through an RNA intermediate by a reverse transcriptase, are classified into the kingdom Pararnavirae in the realm Riboviria.

Parvoviruses are a family of animal viruses that constitute the family Parvoviridae. They have linear, single-stranded DNA (ssDNA) genomes that typically contain two genes encoding for a replication initiator protein, called NS1, and the protein the viral capsid is made of. The coding portion of the genome is flanked by telomeres at each end that form into hairpin loops that are important during replication. Parvovirus virions are small compared to most viruses, at 23–28 nanometers in diameter, and contain the genome enclosed in an icosahedral capsid that has a rugged surface.

Escherichia virus T4 is a species of bacteriophages that infect Escherichia coli bacteria. It is a double-stranded DNA virus in the subfamily Tevenvirinae from the family Myoviridae. T4 is capable of undergoing only a lytic lifecycle and not the lysogenic lifecycle. The species was formerly named T-even bacteriophage, a name which also encompasses, among other strains, Enterobacteria phage T2, Enterobacteria phage T4 and Enterobacteria phage T6.

Caudoviricetes is an class of viruses known as the tailed bacteriophages. Under the Baltimore classification scheme, the Caudoviricetes are group I viruses as they have double stranded DNA (dsDNA) genomes, which can be anywhere from 18,000 base pairs to 500,000 base pairs in length. The virus particles have a distinct shape; each virion has an icosahedral head that contains the viral genome, and is attached to a flexible tail by a connector protein. The order encompasses a wide range of viruses, many containing genes of similar nucleotide sequence and function. However, some tailed bacteriophage genomes can vary quite significantly in nucleotide sequence, even among the same genus. Due to their characteristic structure and possession of potentially homologous genes, it is believed these bacteriophages possess a common origin.

Herpesviridae is a large family of DNA viruses that cause infections and certain diseases in animals, including humans. The members of this family are also known as herpesviruses. The family name is derived from the Greek word ἕρπειν, referring to spreading cutaneous lesions, usually involving blisters, seen in flares of herpes simplex 1, herpes simplex 2 and herpes zoster (shingles). In 1971, the International Committee on the Taxonomy of Viruses (ICTV) established Herpesvirus as a genus with 23 viruses among four groups. As of 2020, 115 species are recognized, all but one of which are in one of the three subfamilies. Herpesviruses can cause both latent and lytic infections.
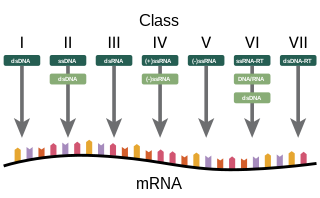
Baltimore classification is a system used to classify viruses based on their manner of messenger RNA (mRNA) synthesis. By organizing viruses based on their manner of mRNA production, it is possible to study viruses that behave similarly as a distinct group. Seven Baltimore groups are described that take into consideration whether the viral genome is made of deoxyribonucleic acid (DNA) or ribonucleic acid (RNA), whether the genome is single- or double-stranded, and whether the sense of a single-stranded RNA genome is positive or negative.

Herpes simplex virus1 and 2, also known by their taxonomical names Human alphaherpesvirus 1 and Human alphaherpesvirus 2, are two members of the human Herpesviridae family, a set of viruses that produce viral infections in the majority of humans. Both HSV-1 and HSV-2 are very common and contagious. They can be spread when an infected person begins shedding the virus.

Gammaherpesvirinae is a subfamily of viruses in the order Herpesvirales and in the family Herpesviridae. Viruses in Gammaherpesvirinae are distinguished by reproducing at a more variable rate than other subfamilies of Herpesviridae. Mammals serve as natural hosts. There are 43 species in this subfamily, divided among 7 genera with three species unassigned to a genus. Diseases associated with this subfamily include: HHV-4: infectious mononucleosis. HHV-8: Kaposi's sarcoma.
Salmonella virus P22 is a bacteriophage in the Podoviridae family that infects Salmonella typhimurium. Like many phages, it has been used in molecular biology to induce mutations in cultured bacteria and to introduce foreign genetic material. P22 has been used in generalized transduction and is an important tool for investigating Salmonella genetics.
HHV Capsid Portal Protein, or HSV-1 UL-6 protein, is the protein which forms a cylindrical portal in the capsid of Herpes simplex virus (HSV-1). The protein is commonly referred to as the HSV-1 UL-6 protein because it is the transcription product of Herpes gene UL-6.
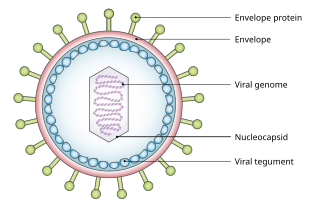
A viral tegument or tegument, more commonly known as a viral matrix, is a cluster of proteins that lines the space between the envelope and nucleocapsid of all herpesviruses. The tegument generally contains proteins that aid in viral DNA replication and evasion of the immune response, typically with inhibition of signalling in the immune system and activation of interferons. The tegument is usually released shortly after infection into the cytoplasm. These proteins are usually formed within the late phase of the viral infectious cycle, after viral genes have been replicated. Much information regarding viral teguments has been gathered from studying herpes simplex virus.

A virus is a submicroscopic infectious agent that replicates only inside the living cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea. Since Dmitri Ivanovsky's 1892 article describing a non-bacterial pathogen infecting tobacco plants and the discovery of the tobacco mosaic virus by Martinus Beijerinck in 1898, more than 9,000 of the millions of virus species have been described in detail. Viruses are found in almost every ecosystem on Earth and are the most numerous type of biological entity. The study of viruses is known as virology, a subspeciality of microbiology.

The Herpesvirales is an order of dsDNA viruses with animal hosts, characterised by a common morphology consisting of an icosahedral capsid enclosed in a glycoprotein-containing lipid envelope. Common infections in humans caused by members of this order include cold sores, genital herpes, chickenpox, shingles, and glandular fever. Herpesvirales is the sole order in the class Herviviricetes, which is the sole class in the phylum Peploviricota.
Tristromaviridae is a family of viruses. Archaea of the genera Thermoproteus and Pyrobaculum serve as natural hosts. Tristromaviridae is the sole family in the order Primavirales. There are two genera and three species in the family.
Escherichia virus CC31, formerly known as Enterobacter virus CC31, is a dsDNA bacteriophage of the subfamily Tevenvirinae responsible for infecting the bacteria family of Enterobacteriaceae. It is one of two discovered viruses of the genus Karamvirus, diverging away from the previously discovered T4virus, as a clonal complex (CC). CC31 was first isolated from Escherichia coli B strain S/6/4 and is primarily associated with Escherichia, even though is named after Enterobacter.
Ranid herpesvirus 1 (RaHV-1), also known as the Lucké tumor herpesvirus (LTHV), is a double-stranded DNA virus within the order Herpesvirales. The virus was initially observed within renal tumors in 1934 by Baldwin Lucké, and more recently has become identifiable through the use of PCR in samples isolated from frog tumors. RaHV-1 causes renal tumors within the northern leopard frog, Rana pipiens. The virus has not yet been isolated in vitro within cell lines, meaning that while its existence and symptoms are fairly evident, its methods of transmission, cell infection, and reproduction are largely unknown.
In virology, realm is the highest taxonomic rank established for viruses by the International Committee on Taxonomy of Viruses (ICTV), which oversees virus taxonomy. Six virus realms are recognized and united by specific highly conserved traits:

Varidnaviria is a realm of viruses that includes all DNA viruses that encode major capsid proteins that contain a vertical jelly roll fold. The major capsid proteins (MCP) form into pseudohexameric subunits of the viral capsid, which stores the viral deoxyribonucleic acid (DNA), and are perpendicular, or vertical, to the surface of the capsid. Apart from this, viruses in the realm also share many other characteristics, such as minor capsid proteins (mCP) with the vertical jelly roll fold, an ATPase that packages viral DNA into the capsid, and a DNA polymerase that replicates the viral genome.

An archaeal virus is a virus that infects and replicates in archaea, a domain of unicellular, prokaryotic organisms. Archaeal viruses, like their hosts, are found worldwide, including in extreme environments inhospitable to most life such as acidic hot springs, highly saline bodies of water, and at the bottom of the ocean. They have been also found in the human body. The first known archaeal virus was described in 1974 and since then, a large diversity of archaeal viruses have been discovered, many possessing unique characteristics not found in other viruses. Little is known about their biological processes, such as how they replicate, but they are believed to have many independent origins, some of which likely predate the last archaeal common ancestor (LACA).

Adnaviria is a realm of viruses that includes archaeal viruses that have a filamentous virion and a linear, double-stranded DNA genome. The genome exists in A-form (A-DNA) and encodes a dimeric major capsid protein (MCP) that contains the SIRV2 fold, a type of alpha-helix bundle containing four helices. The virion consists of the genome encased in capsid proteins to form a helical nucleoprotein complex. For some viruses, this helix is surrounded by a lipid membrane called an envelope. Some contain an additional protein layer between the nucleoprotein helix and the envelope. Complete virions are long and thin and may be flexible or a stiff like a rod.















