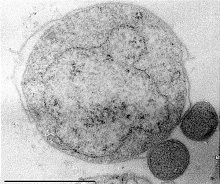
Nanoarchaeum equitans is a species of marine archaea that was discovered in 2002 in a hydrothermal vent off the coast of Iceland on the Kolbeinsey Ridge by Karl Stetter. It has been proposed as the first species in a new phylum, and is the only species within the genus Nanoarchaeum. Strains of this microbe were also found on the Sub-polar Mid Oceanic Ridge, and in the Obsidian Pool in Yellowstone National Park. Since it grows in temperatures approaching boiling, at about 80 °C (176 °F), it is considered to be a thermophile. It grows best in environments with a pH of 6, and a salinity concentration of 2%. Nanoarchaeum appears to be an obligate symbiont on the archaeon Ignicoccus; it must be in contact with the host organism to survive. Nanoarchaeum equitans cannot synthesize lipids but obtains them from its host. Its cells are only 400 nm in diameter, making it the smallest known living organism, and the smallest known archaeon.

Nanoarchaeota is a proposed phylum in the domain Archaea that currently has only one representative, Nanoarchaeum equitans, which was discovered in a submarine hydrothermal vent and first described in 2002.

The Korarchaeota is a proposed phylum within the Archaea. The name is derived from the Greek noun koros or kore, meaning young man or young woman, and the Greek adjective archaios which means ancient. They are also known as Xenarchaeota. The name is equivalent to Candidatus Korarchaeota, and they go by the name Xenarchaeota or Xenarchaea as well.
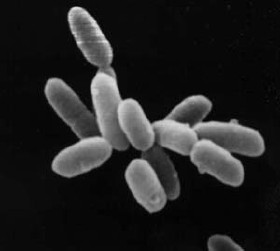
Euryarchaeota is a kingdom of archaea. Euryarchaeota are highly diverse and include methanogens, which produce methane and are often found in intestines; halobacteria, which survive extreme concentrations of salt; and some extremely thermophilic aerobes and anaerobes, which generally live at temperatures between 41 and 122 °C. They are separated from the other archaeans based mainly on rRNA sequences and their unique DNA polymerase.

Metagenomics is the study of genetic material recovered directly from environmental or clinical samples by a method called sequencing. The broad field may also be referred to as environmental genomics, ecogenomics, community genomics or microbiomics.
Archaeoglobus is a genus of the phylum Euryarchaeota. Archaeoglobus can be found in high-temperature oil fields where they may contribute to oil field souring.

Ferroplasma is a genus of Archaea that belong to the family Ferroplasmaceae. Members of the Ferroplasma are typically acidophillic, pleomorphic, irregularly shaped cocci.
Pyrobaculum is a genus of the Thermoproteaceae.

Archaea is a domain of single-celled organisms. These microorganisms lack cell nuclei and are therefore prokaryotic. Archaea were initially classified as bacteria, receiving the name archaebacteria, but this term has fallen out of use.

The Nitrososphaerota are a phylum of the Archaea proposed in 2008 after the genome of Cenarchaeum symbiosum was sequenced and found to differ significantly from other members of the hyperthermophilic phylum Thermoproteota. Three described species in addition to C. symbiosum are Nitrosopumilus maritimus, Nitrososphaera viennensis, and Nitrososphaera gargensis. The phylum was proposed in 2008 based on phylogenetic data, such as the sequences of these organisms' ribosomal RNA genes, and the presence of a form of type I topoisomerase that was previously thought to be unique to the eukaryotes. This assignment was confirmed by further analysis published in 2010 that examined the genomes of the ammonia-oxidizing archaea Nitrosopumilus maritimus and Nitrososphaera gargensis, concluding that these species form a distinct lineage that includes Cenarchaeum symbiosum. The lipid crenarchaeol has been found only in Nitrososphaerota, making it a potential biomarker for the phylum. Most organisms of this lineage thus far identified are chemolithoautotrophic ammonia-oxidizers and may play important roles in biogeochemical cycles, such as the nitrogen cycle and the carbon cycle. Metagenomic sequencing indicates that they constitute ~1% of the sea surface metagenome across many sites.
Nanohaloarchaea is a clade of diminutive archaea with small genomes and limited metabolic capabilities, belonging to the DPANN archaea. They are ubiquitous in hypersaline habitats, which they share with the extremely halophilic haloarchaea.
Nitrososphaera is a mesophilic genus of ammonia-oxidizing Crenarchaeota. The first Nitrososphaera organism was discovered in garden soils at the University of Vienna leading to the categorization of a new genus, family, order and class of Archaea. This genus is contains three distinct species: N. viennensis, Ca. N. gargensis, and Ca N. evergladensis. Nitrososphaera are chemolithoautotrophs and have important biogeochemical roles as nitrifying organisms.
The "Aigarchaeota" are a proposed archaeal phylum of which the main representative is Caldiarchaeum subterraneum. It is not yet clear if this represents a new phylum or a Nitrososphaerota order, since the genome of Caldiarchaeum subterraneum encodes several Nitrososphaerota-like features. The name "Aigarchaeota" comes from the Greek αυγή, avgí, meaning "dawn" or "aurora", for the intermediate features of hyperthermophilic and mesophilic life during the evolution of its lineage.
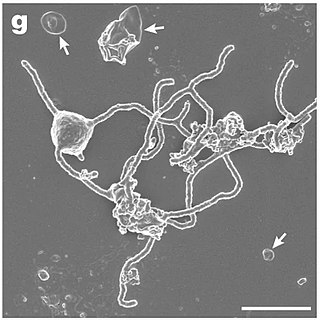
Lokiarchaeota is a proposed phylum of the Archaea. The phylum includes all members of the group previously named Deep Sea Archaeal Group, also known as Marine Benthic Group B. Lokiarchaeota is part of the superphylum Asgard containing the phyla: Lokiarchaeota, Thorarchaeota, Odinarchaeota, Heimdallarchaeota, and Helarchaeota. A phylogenetic analysis disclosed a monophyletic grouping of the Lokiarchaeota with the eukaryotes. The analysis revealed several genes with cell membrane-related functions. The presence of such genes support the hypothesis of an archaeal host for the emergence of the eukaryotes; the eocyte-like scenarios.

Parvarchaeota is a phylum of archaea belonging to the DPANN archaea. They have been discovered in acid mine drainage waters and later in marine sediments. The cells of these organisms are extremely small consistent with small genomes. Metagenomic techniques allow obtaining genomic sequences from non-cultured organisms, which were applied to determine this phylum.
Hadesarchaea, formerly called the South-African Gold Mine Miscellaneous Euryarchaeal Group, are a class of thermophile microorganisms that have been found in deep mines, hot springs, marine sediments, and other subterranean environments.
Nitrososphaera gargensis is a non-pathogenic, small coccus measuring 0.9 ± 0.3 μm in diameter. N. gargensis is observed in small abnormal cocci groupings and uses its archaella to move via chemotaxis. Being an Archaeon, Nitrososphaera gargensis has a cell membrane composed of crenarchaeol, its isomer, and a distinct glycerol dialkyl glycerol tetraether (GDGT), which is significant in identifying ammonia-oxidizing archaea (AOA). The organism plays a role in influencing ocean communities and food production.

DPANN is a superphylum of Archaea first proposed in 2013. Many members show novel signs of horizontal gene transfer from other domains of life. They are known as nanoarchaea or ultra-small archaea due to their smaller size (nanometric) compared to other archaea.
"Candidatus Thorarchaeota", or simply Thorarchaeota, is a phylum within the superphylum Asgard archaea. The Asgard superphylum represents the closest prokaryotic relatives of eukaryotes. Since there is such a close relation between the two different domains, it provides further evidence to the two-domain tree of life theory which states that eukaryotes branched from the archaeal domain. Asgard archaea are single cell marine microbes that contain branch like appendages and have genes that are similar to eukarya. The asgard archaea superphylum is composed of Thorarchaeota, Lokiarchaeota, Odinarchaeota, and Heimdallarchaeota. Thorarchaeota were first identified from the sulfate-methane transition zone in tidewater sediments. Thorarcheota are widely distributed in marine and freshwater sediments.
TM7x, also known as Nanosynbacter lyticus type strain TM7x HMT 952. is a phylotype of one of the most enigmatic phyla, Candidatus Saccharibacteria, formerly candidate phylum TM7. It is the only member of the candidate phylum that has been cultivated successfully from the human oral cavity, and stably maintained in vitro. and serves as a crucial paradigm. of the newly described Candidate Phyla Radiation (CPR). The cultivated oral taxon is designated as Saccharibacteria oral taxon TM7x. TM7x has a unique lifestyle in comparison to other bacteria that are associated with humans. It is an obligate epibiont parasite, or an "epiparasite", growing on the surface of its host bacterial species Actinomyces odontolyticus subspecies actinosynbacter strain XH001, which is referred to as the "basibiont". Actinomyces species are one of the early microbial colonizers in the oral cavity. Together, they exhibit parasitic epibiont symbiosis.










